Physical Map
Definition
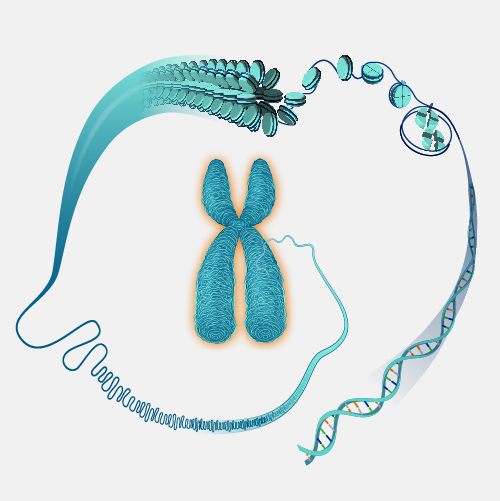
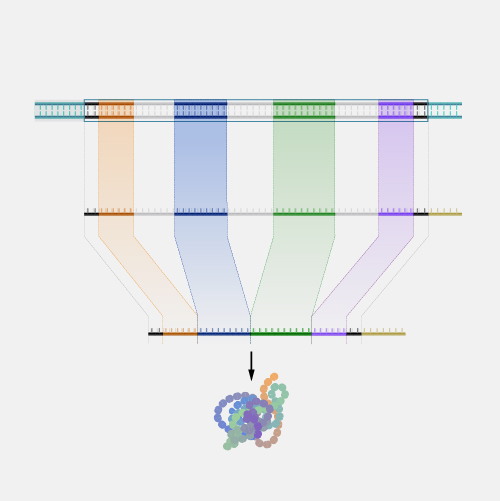
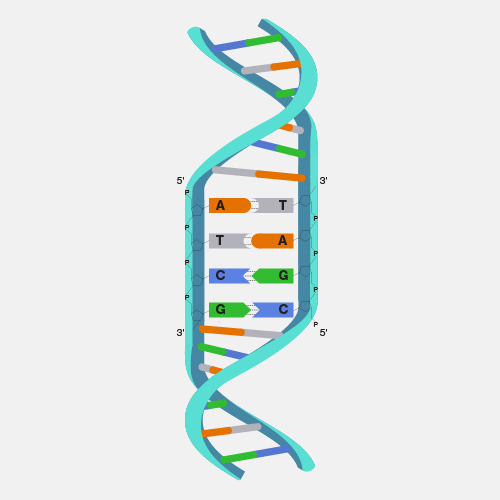
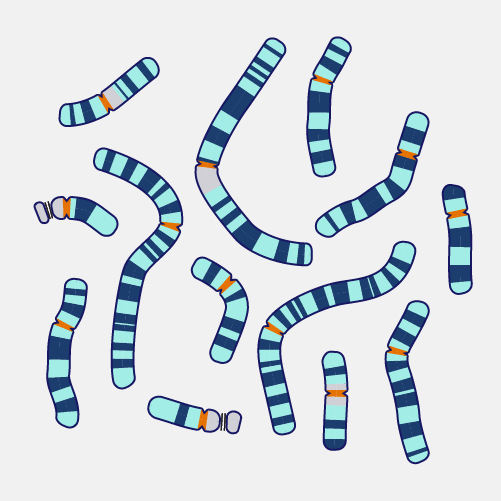
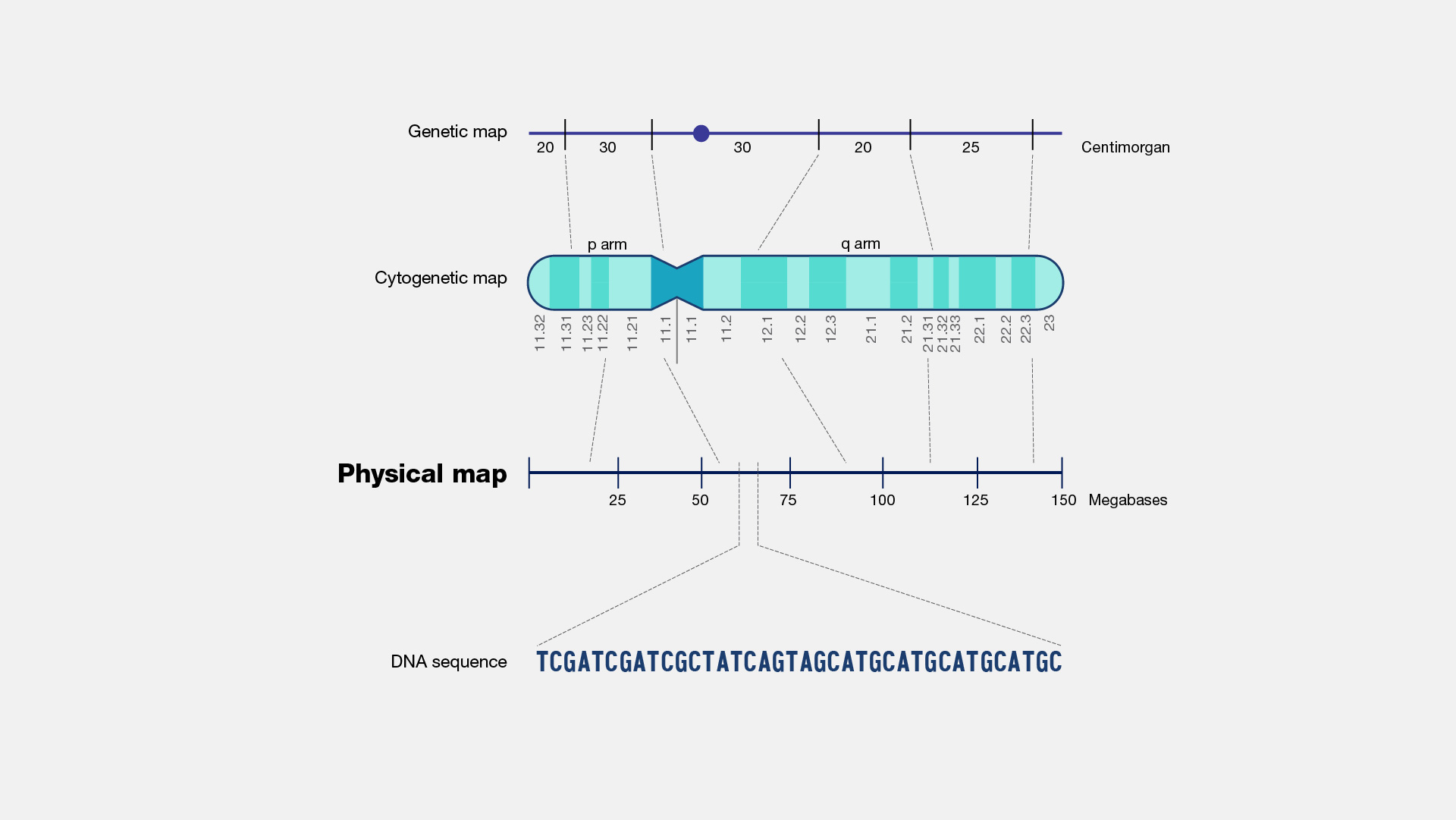
A physical map, as related to genomics, is a graphical representation of physical locations of landmarks or markers (such as genes, variants and other DNA sequences of interest) within a chromosome or genome. A complete genome sequence is one type of physical map. Physical maps are used to identify genes or other sequences believed to play a role in health conditions or diseases. They are also valuable in providing an organizational framework for generating complete sequences of genomes.
Narration
Now that genomes can be sequenced quite efficiently and inexpensively, genomicists do not talk as often about physical maps. But previously, it was very important to first develop a map of a segment of DNA of interest or a genome that you wanted to study prior to actually starting to sequence -- or read-- that DNA. So, during the Human Genome Project, researchers first developed very high-quality physical maps of each human chromosome, and then proceeded to use those maps as an organizational framework for determining the sequence of each chromosome. One analogy that can be useful to consider is prior to reading a book, you want to make sure you have all the book’s pages in the correct order. So, you can consider the number on each page of a book to being a type of physical map of that book.