Shotgun Sequencing
Definition
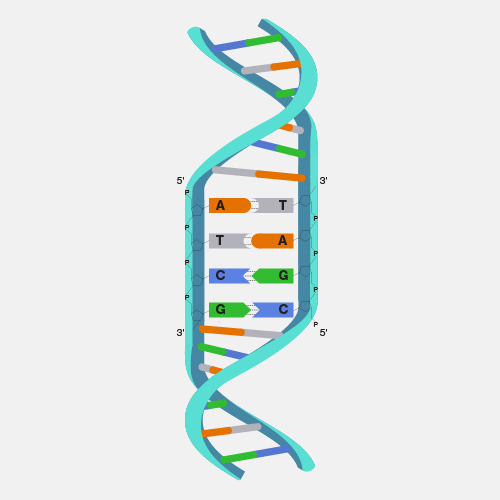
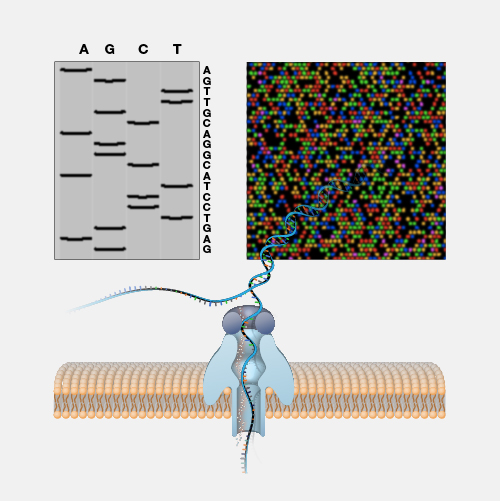
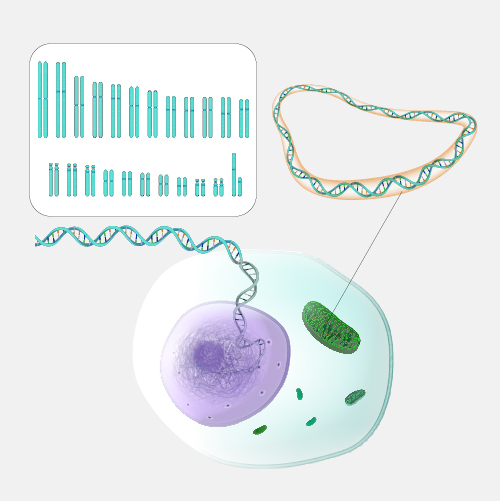
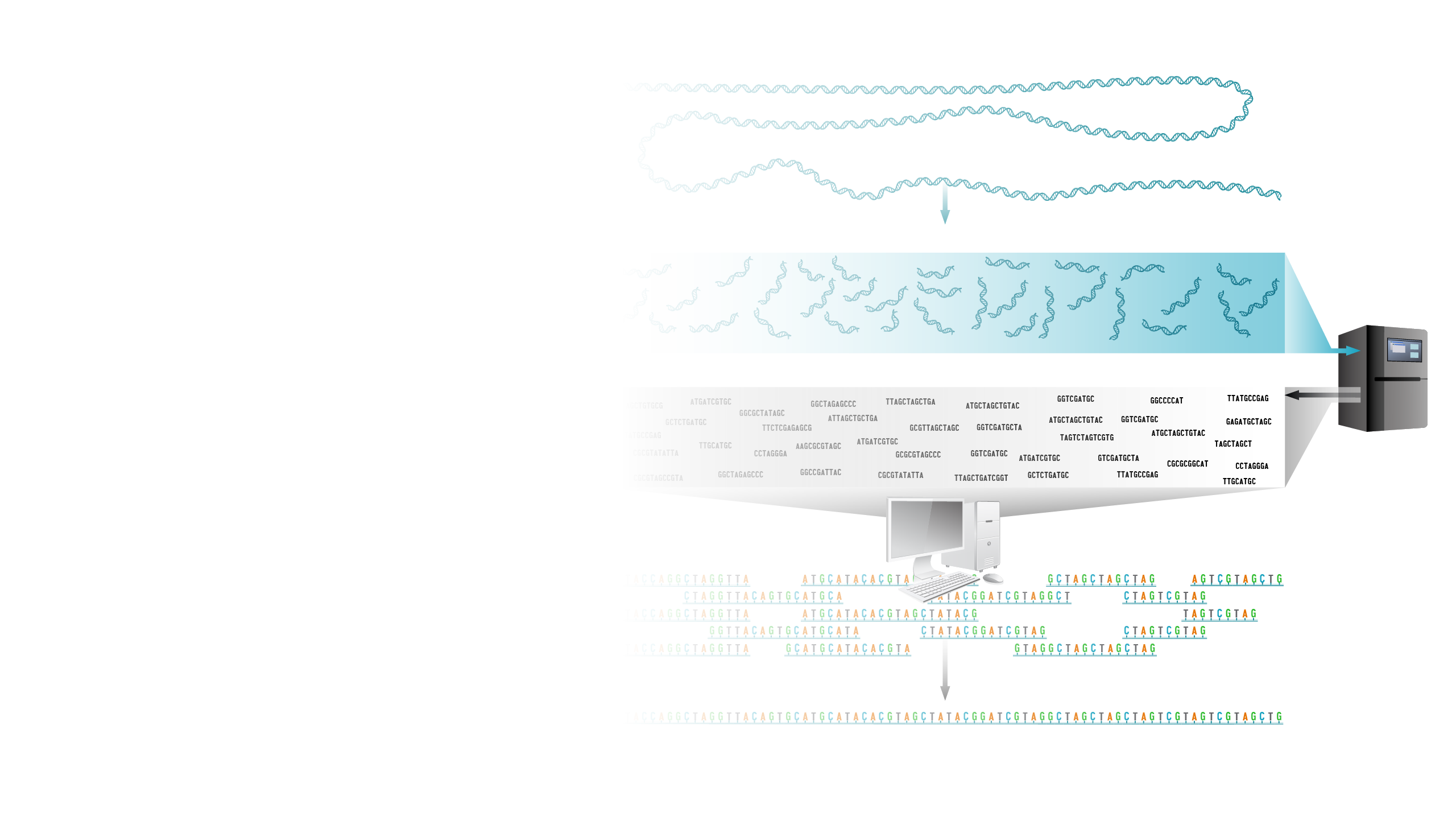
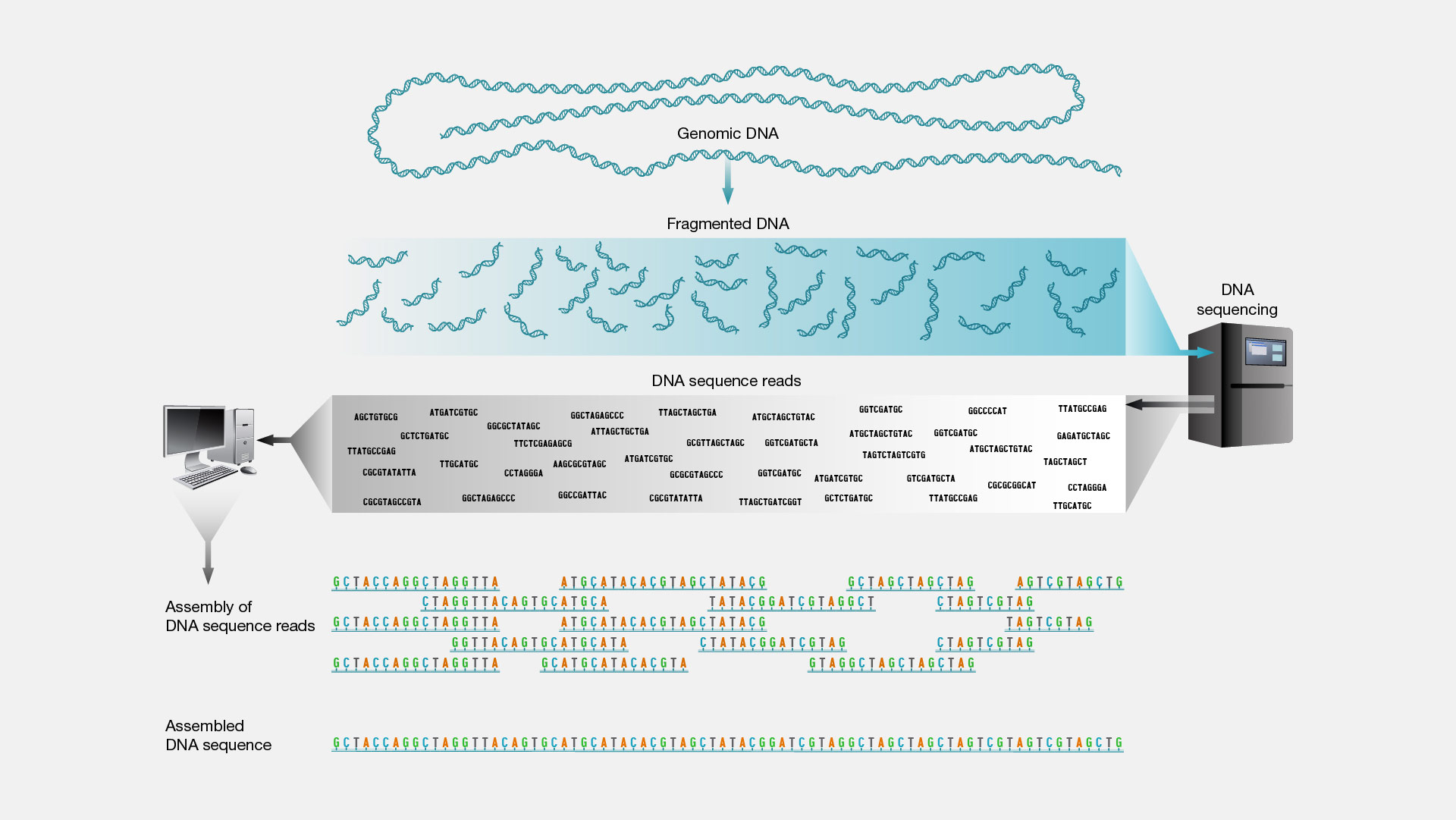
Shotgun sequencing is a laboratory technique for determining the DNA sequence of an organism’s genome. The method involves randomly breaking up the genome into small DNA fragments that are sequenced individually. A computer program looks for overlaps in the DNA sequences, using them to reassemble the fragments in their correct order to reconstitute the genome.
Narration
Imagine taking a page in a book, making hundreds of copies of it, taking those copies and putting them in a paper shredder that magically creates strips of paper that contain various bits and pieces of the original sentences on the starting page. Then imagine reading each of those strips containing the bits and pieces of the sentences, looking for overlapping words and phrases, and eventually being able to reconstruct the entire text on the starting page by having read enough of those shreds of paper. In essence, that is shotgun DNA sequencing. Instead of a page of a book, a scientist starts with a piece of clone DNA or even an entire genome. The DNA is broken apart and many many many many many sequence reads are generated from the DNA pieces. All of those data are analyzed by a computer program that looks for overlapping stretches of sequence and eventually puts that DNA puzzle back together.