Antisense
Definition
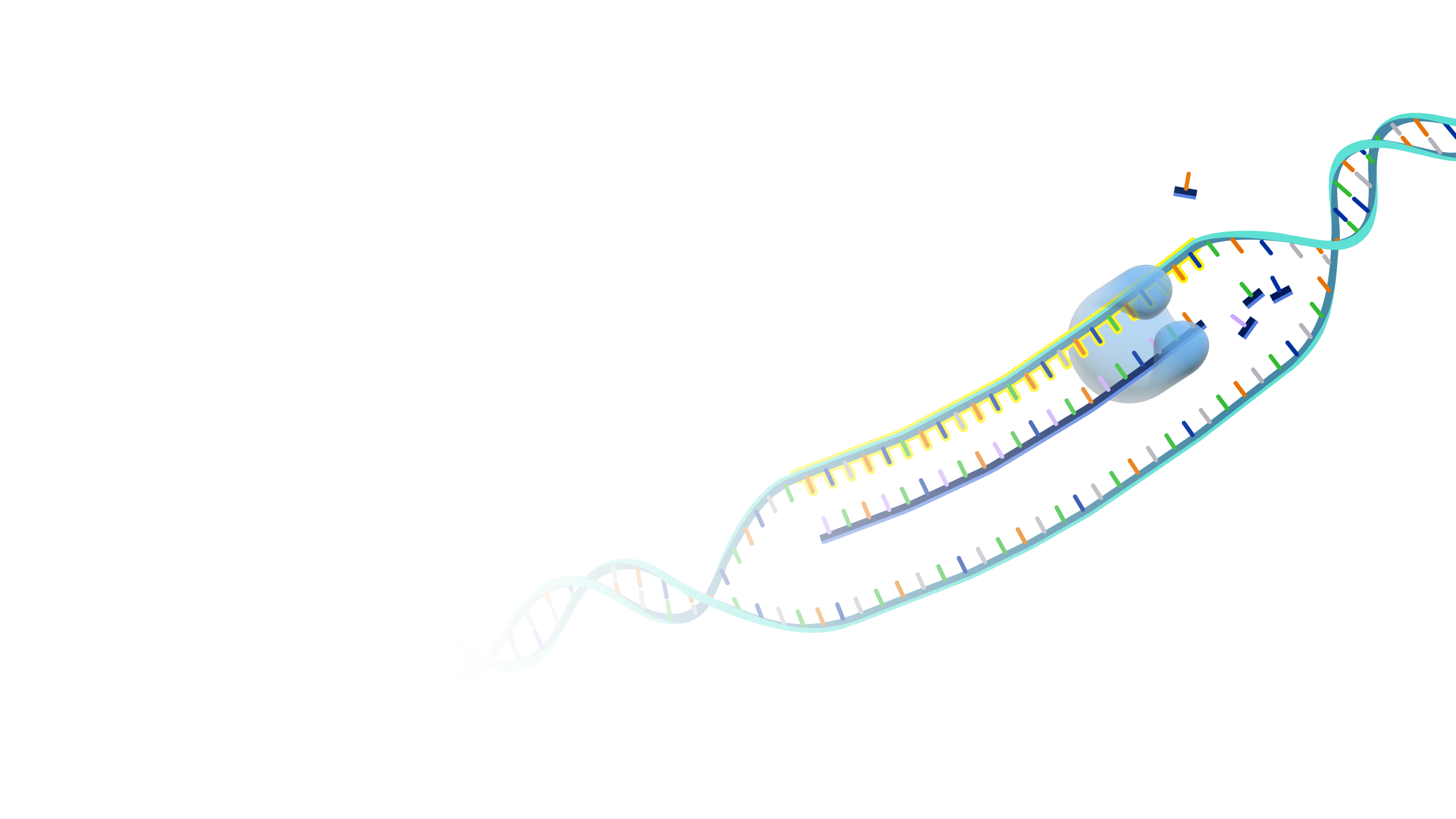
Antisense is the non-coding DNA strand of a gene. In a cell, antisense DNA serves as the template for producing messenger RNA (mRNA), which directs the synthesis of a protein.
Narration
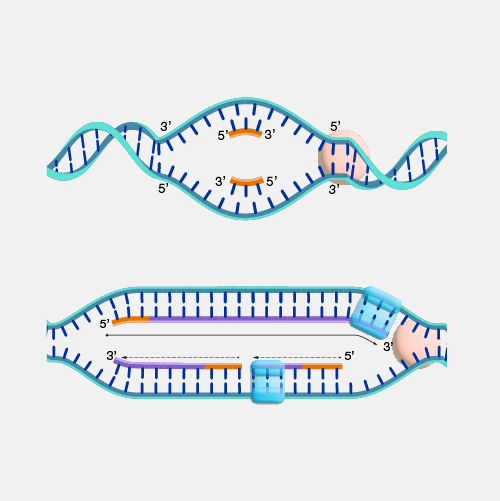
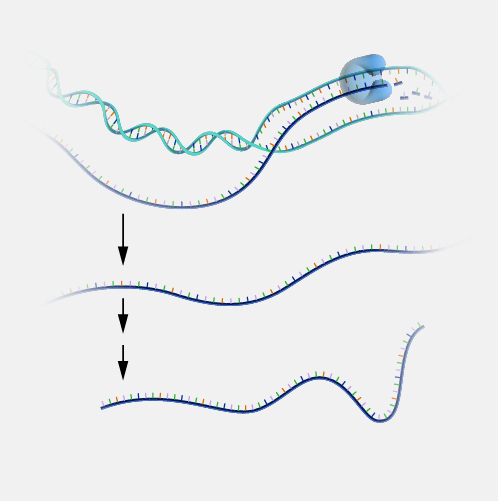
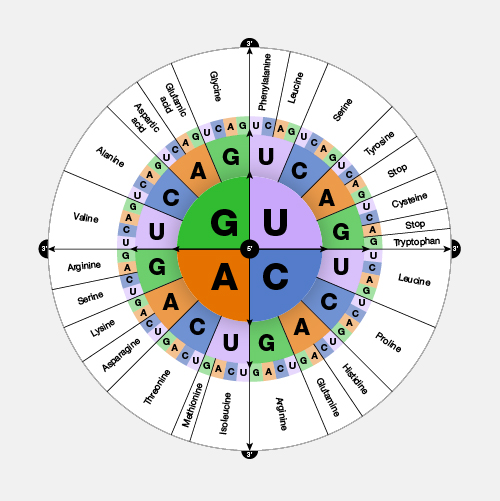
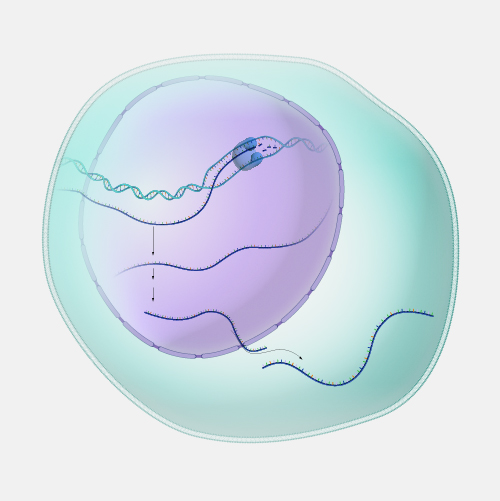
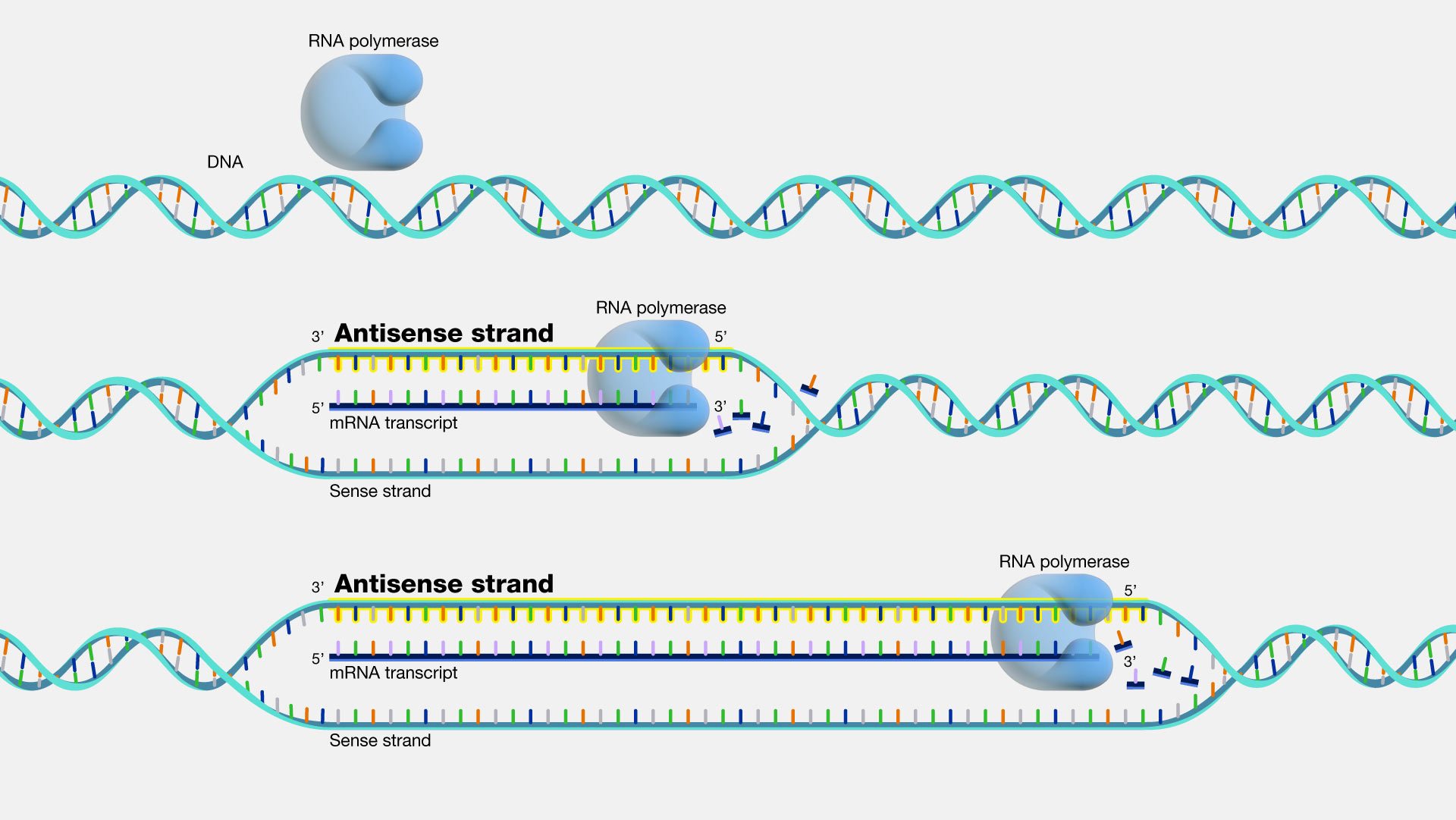
Antisense. DNA is a two-stranded molecule. When the DNA is read so that it can ultimately be translated into a protein, it can only be read in one direction. One strand of DNA is called the sense strand because when you read it in the right direction it provides the code to make a protein. In two-stranded DNA, the sense strand is bonded to an opposite DNA strand that is called the antisense or noncoding strand. Each of the two DNA strands is made up of a series of bases that are often labeled with the letters A, T, C, and G. Every base on the sense strand naturally pairs with a complementary or matching base on the antisense strand. So A on the sense strand always pairs with a T on the antisense strand and C always pairs with G. During protein productions, the two DNA strands temporarily separate. The antisense strand then becomes a template for making messenger RNA, and the resulting messenger RNA is then a sense messenger RNA that, essentially, matches the sense DNA strand. The messenger RNA is then involved in coding for the corresponding protein.