DNA Microarray Technology Fact Sheet
The DNA microarray is a tool used to determine whether the DNA from a particular individual contains a mutation in genes.
What is a DNA microarray?
Scientists know that a mutation - or alteration - in a particular gene's DNA may contribute to a certain disease. However, it can be very difficult to develop a test to detect these mutations, because most large genes have many regions where mutations can occur. For example, researchers believe that mutations in the genes BRCA1 and BRCA2 cause as many as 60 percent of all cases of hereditary breast and ovarian cancers. But there is not one specific mutation responsible for all of these cases. Researchers have already discovered over 800 different mutations in BRCA1 alone.The DNA microarray is a tool used to determine whether the DNA from a particular individual contains a mutation in genes like BRCA1 and BRCA2. The chip consists of a small glass plate encased in plastic. Some companies manufacture microarrays using methods similar to those used to make computer microchips. On the surface, each chip contains thousands of short, synthetic, single-stranded DNA sequences, which together add up to the normal gene in question, and to variants (mutations) of that gene that have been found in the human population.
-
What is a DNA microarray?
Scientists know that a mutation - or alteration - in a particular gene's DNA may contribute to a certain disease. However, it can be very difficult to develop a test to detect these mutations, because most large genes have many regions where mutations can occur. For example, researchers believe that mutations in the genes BRCA1 and BRCA2 cause as many as 60 percent of all cases of hereditary breast and ovarian cancers. But there is not one specific mutation responsible for all of these cases. Researchers have already discovered over 800 different mutations in BRCA1 alone.The DNA microarray is a tool used to determine whether the DNA from a particular individual contains a mutation in genes like BRCA1 and BRCA2. The chip consists of a small glass plate encased in plastic. Some companies manufacture microarrays using methods similar to those used to make computer microchips. On the surface, each chip contains thousands of short, synthetic, single-stranded DNA sequences, which together add up to the normal gene in question, and to variants (mutations) of that gene that have been found in the human population.

What is a DNA microarray used for?
When they were first introduced, DNA microarrays were used only as a research tool. Scientists continue today to conduct large-scale population studies - for example, to determine how often individuals with a particular mutation actually develop breast cancer, or to identify the changes in gene sequences that are most often associated with particular diseases. This has become possible because, just as is the case for computer chips, very large numbers of 'features' can be put on microarray chips, representing a very large portion of the human genome.
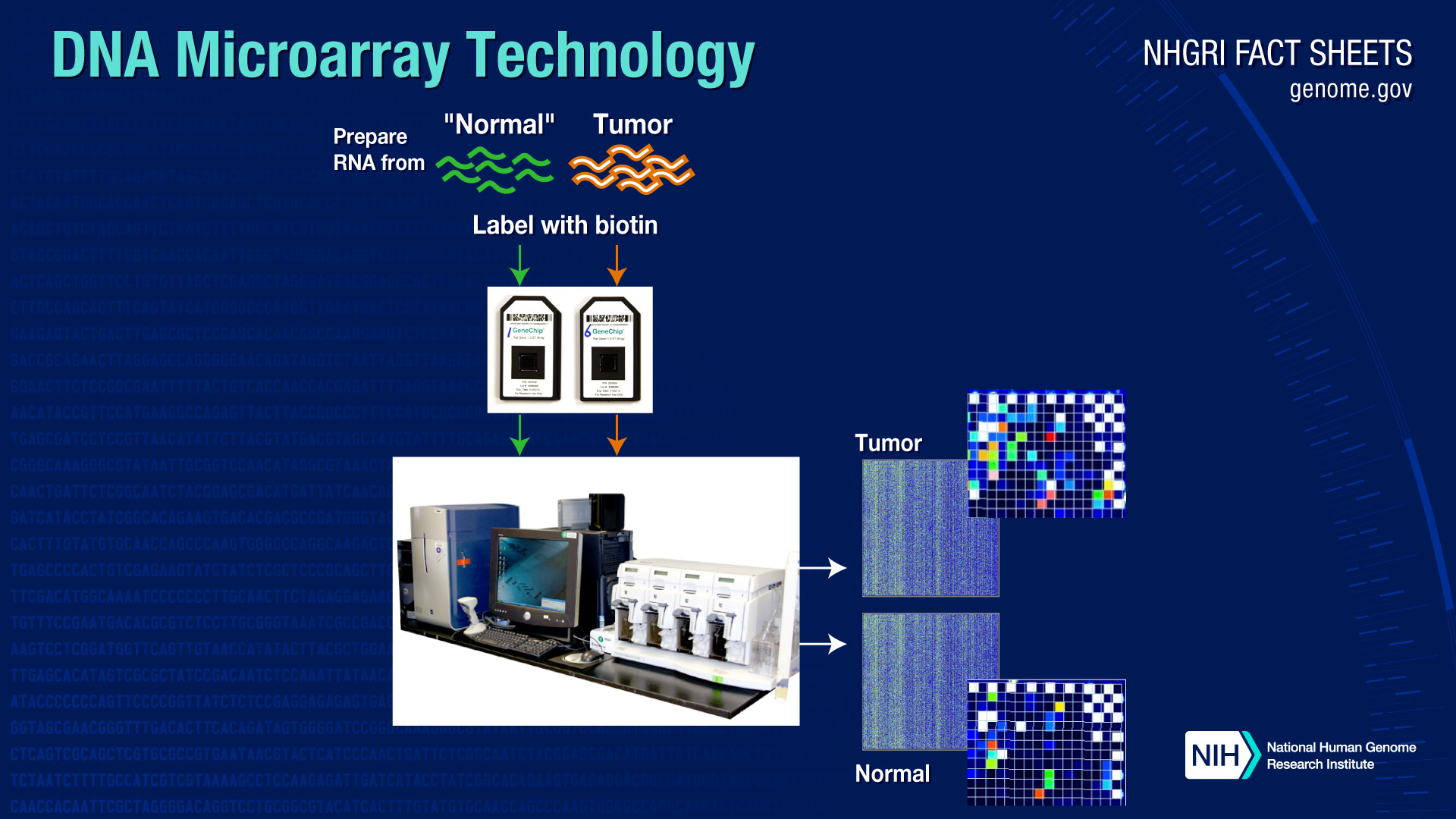
Microarrays can also be used to study the extent to which certain genes are turned on or off in cells and tissues. In this case, instead of isolating DNA from the samples, RNA (which is a transcript of the DNA) is isolated and measured.
Today, DNA microarrays are used in clinical diagnostic tests for some diseases. Sometimes they are also used to determine which drugs might be best prescribed for particular individuals, because genes determine how our bodies handle the chemistry related to those drugs.With the advent of new DNA sequencing technologies, some of the tests for which microarrays were used in the past now use DNA sequencing instead. But microarray tests still tend to be less expensive than sequencing, so they may be used for very large studies, as well as for some clinical tests.
-
What is a DNA microarray used for?
When they were first introduced, DNA microarrays were used only as a research tool. Scientists continue today to conduct large-scale population studies - for example, to determine how often individuals with a particular mutation actually develop breast cancer, or to identify the changes in gene sequences that are most often associated with particular diseases. This has become possible because, just as is the case for computer chips, very large numbers of 'features' can be put on microarray chips, representing a very large portion of the human genome.
Microarrays can also be used to study the extent to which certain genes are turned on or off in cells and tissues. In this case, instead of isolating DNA from the samples, RNA (which is a transcript of the DNA) is isolated and measured.
Today, DNA microarrays are used in clinical diagnostic tests for some diseases. Sometimes they are also used to determine which drugs might be best prescribed for particular individuals, because genes determine how our bodies handle the chemistry related to those drugs.With the advent of new DNA sequencing technologies, some of the tests for which microarrays were used in the past now use DNA sequencing instead. But microarray tests still tend to be less expensive than sequencing, so they may be used for very large studies, as well as for some clinical tests.
How does a DNA microarray work?
To determine whether an individual possesses a mutation for a particular disease, a scientist first obtains a sample of DNA from the patient's blood as well as a control sample - one that does not contain a mutation in the gene of interest.
The researcher then denatures the DNA in the samples - a process that separates the two complementary strands of DNA into single-stranded molecules. The next step is to cut the long strands of DNA into smaller, more manageable fragments and then to label each fragment by attaching a fluorescent dye (there are other ways to do this, but this is one common method). The individual's DNA is labeled with green dye and the control - or normal - DNA is labeled with red dye. Both sets of labeled DNA are then inserted into the chip and allowed to hybridize - or bind - to the synthetic DNA on the chip.
If the individual does not have a mutation for the gene, both the red and green samples will bind to the sequences on the chip that represent the sequence without the mutation (the "normal" sequence).
If the individual does possess a mutation, the individual's DNA will not bind properly to the DNA sequences on the chip that represent the "normal" sequence but instead will bind to the sequence on the chip that represents the mutated DNA.
-
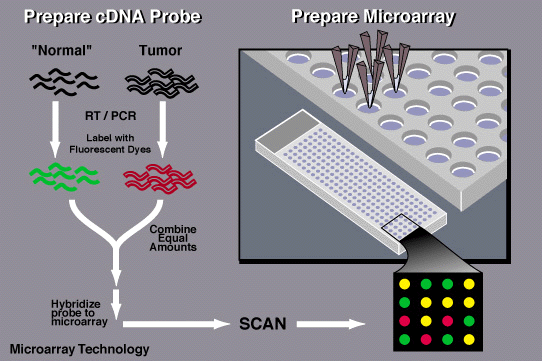
How does a DNA microarray work?
To determine whether an individual possesses a mutation for a particular disease, a scientist first obtains a sample of DNA from the patient's blood as well as a control sample - one that does not contain a mutation in the gene of interest.
The researcher then denatures the DNA in the samples - a process that separates the two complementary strands of DNA into single-stranded molecules. The next step is to cut the long strands of DNA into smaller, more manageable fragments and then to label each fragment by attaching a fluorescent dye (there are other ways to do this, but this is one common method). The individual's DNA is labeled with green dye and the control - or normal - DNA is labeled with red dye. Both sets of labeled DNA are then inserted into the chip and allowed to hybridize - or bind - to the synthetic DNA on the chip.
If the individual does not have a mutation for the gene, both the red and green samples will bind to the sequences on the chip that represent the sequence without the mutation (the "normal" sequence).
If the individual does possess a mutation, the individual's DNA will not bind properly to the DNA sequences on the chip that represent the "normal" sequence but instead will bind to the sequence on the chip that represents the mutated DNA.

Last updated: August 15, 2020
