Last updated: April 04, 2012
NHGRI researchers identify gene for inherited bleeding disorder
NHGRI researchers identify gene for inherited bleeding disorder
By Raymond MacDougall
Associate Communications Director for Intramural Research
 |
The study, published in the July 17, 2011, early online issue of Nature Genetics, is the second publication from the group in the span of a year, cracking the DNA code for the condition after almost a decade of research and international collaboration to locate patients with the disorder. "Finding the GPS gene is an important step in improving our understanding of how certain organelles are formed," said lead author Meral Gunay-Aygun, M.D., NHGRI staff clinician and an associate professor in the Department of Pediatrics at Johns Hopkins Medicine in Baltimore.
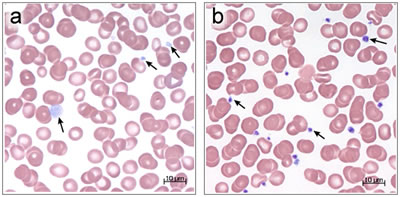
Platelets have two major organelles-alpha-granules and delta-granules. Alpha-granules store proteins required for the formation of blood clots that stop bleeding. GPS is characterized by the absence of alpha-granules and their contents, which produces a bleeding disorder in patients with this condition. The syndrome derives its name from the gray appearance of platelets in light microscope images due to the lack of alpha-granules that give the platelets their normal purple color.
GPS is a progressive condition that is usually detected when patients experience bleeding and low platelet counts. The disorder also causes scar tissue in the bone marrow and enlargement of the spleen. Bleeding tendency in GPS is usually mild to moderate, but deaths have occurred due to hemorrhages, especially in advanced stages of the disease when platelet counts decrease even more.
Dr. Gunay-Aygun, lead investigator of the study, launched an NIH protocol in 2004 and recruited 26 GPS patients from 15 families around the world. The first attempts to locate the gene for GPS involved genetic linkage analysis of the DNA from patients recruited to NIH. The linkage analysis narrowed down the location of the GPS gene to a large region on chromosome 3 that was shared by the affected individuals from all families. Last year, the group published a paper in the journal Blood that reported the mapping results and described in detail the clinical features and natural history of the disease.
The 2010 study, based upon data from 25 patients in 14 families, detailed efforts to map the region including the GPS gene, which had been narrowed down to a region of 197 protein-coding genes on chromosome 3. In the current publication, the researchers zeroed in on the NBEAL2 gene through whole-exome sequencing, an approach that decodes the 1-2 percent of the genome that contains protein-coding genes. The sequencing and analysis was performed at the NIH Intramural Sequencing Center (NISC), administered by NHGRI.
"During the course of the project, we adapted our search to available technologies, getting progressively closer to finding the gene. Whole-exome sequencing enabled us to exclude a large number of genes and eventually identify NBEAL2 as the GPS gene." Dr. Gunay-Aygun is a member of NHGRI's Medical Genetics Branch, where a key research focus is on disorders characterized by abnormalities in formation of organelle structures within cells. Impaired organelles, including lysosomes, which perform a housekeeping role within cells, can cause symptoms such as albinism, bleeding, lung damage and neurological problems depending on the particular disorder.
"Even though it is relatively rare, GPS is a well-known disorder among hematologists because the basic defect is so clear-cut," Dr. Gunay-Aygun said about the disease first described in 1971. "Alpha-granules, the most abundant organelles in platelets are missing. What we learn about the function of NBEAL2 protein in platelets will probably help us understand how organelles form in other cells types as well."
Despite early enthusiasm for finding the gene responsible for GPS-and long nights sequencing genes in the lab-Dr. Gunay-Aygun's expectations waivered. "This project seemed to run out of luck many times," she said. But, according to Dr. Gunay-Aygun, perseverance paid off.
Initially the team sequenced more than 30 selected candidate genes in the region using traditional DNA sequencing methods but could not find any convincing mutations. Attempts to sequence all the genes in the region at NISC excluded 75 percent of the target region. Subsequent whole-exome sequencing did not identify mutations but significantly decreased the number of genes.
The researchers ultimately identified mutations in NBEAL2 by performing a final round of analysis of the remaining difficult-to-sequence genes. NBEAL2 is a large gene containing 52 exons. "It is not inviting to sequence it manually in the lab," said co-author Marjan Huizing, Ph.D., NHGRI associate investigator. Despite the difficulty, the researchers found mutations in each of the 15 GPS families in the study. "These are different mutations, but they all impair the function of the protein," said Dr. Huizing.
The researchers have only begun to explore the function of the NBEAL2 protein and note that it is similar in composition to the protein defective in Chediak-Higashi syndrome, another organelle disorder. "The Chediak-Higashi syndrome gene was identified more than 10 years ago and its function is still not completely understood," said Dr. Gunay-Aygun. "It sometimes takes a long time to determine the exact function of a protein."
Dr. Gunay-Aygun hopes that the study of the underlying cell biological defect in GPS will contribute to our overall understanding of how organelles form and lead to new strategies for treatment of these disorders.
