Last updated: March 05, 2012
When cellular power plants go bad
Genome Advance of the Month
When cellular power plants go bad
By Jonathan Max Gitlin
Science Policy Analyst
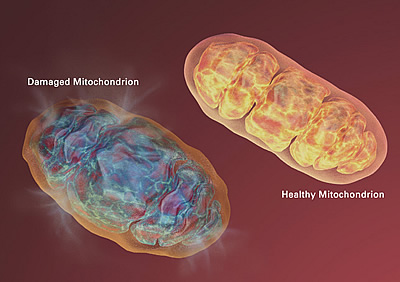
The evolution of this relationship, and the subsequent development of the cell's power plant known as the mitochondria, gave rise to eukaryotes, an entire group of organisms whose cells contain complex structures enclosed in membranes. As a result, all eukaryotic cells have both a nuclear genome — in the case of humans, the 3 billion base pairs we know as the human genome — and a mitochondrial genome, a much simpler, circular genome of around 16,000 base pairs.
This genome advance of the month focuses on the use of DNA sequencing for diagnosing severe and confounding mitochondrial diseases. As many as 4,000 children are born in the United States with mitochondrial dysfunction every year. Researchers have identified 77 genes known to cause mitochondrial diseases, but the culprit gene remains elusive in half of all cases. Even Leigh's Disease, one of the most common mitochondrial diseases that affects cognitive development, can be caused by mutations in more than 25 different genes.
Diagnosing mitochondrial dysfunction is difficult due to a number of factors. The heart, lung, brain and muscles tend to be the most severely affected, with many different cell types or organs involved. As a result, symptoms vary widely among patients, including muscle fibers that do not function (myopathy, a muscular disease), lactic acidosis (when lactic acid builds ups in the bloodstream faster than it can be removed), seizures, ataxia (lack of coordination while performing voluntary movements), peripheral neuropathy (malfunction of the nerves running from the brain and spinal cord to the rest of the body), blindness, deafness, gastrointestinal problems, liver failure, and bone marrow dysfunction. Symptoms can appear in utero (usually resulting in miscarriage or stillbirth), during infancy or in adulthood. The condition may be inherited from one or both parents, or the result of spontaneous mutation, either in the mitochondrial or the nuclear genome.
Now, a team of scientists in the United States and Australia have used high-throughput DNA sequencing to see if such an approach might lend itself for use in diagnosing mitochondrial diseases. The study, by Sarah Calvo, Ph.D., a computational biologist at the Broad Institute of MIT/Harvard, Alison Compton, Ph.D., a molecular geneticist at the Murdoch Childrens Research Institute, Melbourne, Australia, and their colleagues, was published in Science Translational Medicine in January.
Using samples collected from 42 unrelated infants with clinical evidence of mitochondrial disease but without a definitive molecular diagnosis, (i.e. identifying a causative gene), they sequenced deep coverage of both the mitochondrial genome and roughly 1,000 nuclear genes known to code for mitochondrial proteins.
The researchers sifted through this data, looking for gene variants that could be responsible. Since mitochondrial diseases tend to be both rare and devastating, they hypothesized that the variants causing the disease would be rare in the general population. They also speculated that the variants would alter the protein they coded, and would either be a new dominant mutation or recessively inherited from a parent.
Comparing the sample genes with data on the same genes from healthy individuals in existing databases, the researchers discovered that affected individuals had a five-fold increase in genes where both alleles (one of two or more versions of a gene) contained variants known to disrupt protein function, genes that were prioritized for further analysis, along with the genes known to cause mitochondrial diseases.
Among the 42 infants studied, more than half had mutations present in at least one of the prioritized genes.
Of these:
- Ten infants had mutations in nuclear genes previously known to cause mitochondrial disease.
- Twelve infants had mutations in nuclear genes previously unlinked to mitochondrial disease.
- One infant had a large deletion in their mitochondrial genome.
Although the researchers found only potentially causal mutations in half of the affected infants, they hope that several of the newly-identified variants will prove to cause mitochondrial disease.
These findings challenge some of our existing understanding of mitochondrial conditions. Gene variants that had previously been identified as causing mitochondrial diseases are highly penetrant, meaning that the presence of that gene variant almost always corresponds with the disease, and these variants are usually inherited. Dr. Calvo suggested that many of the sporadic cases, where there isn't an existing family history, may instead be controlled by the interaction of genes at more than one location on the genome, with a number of weak genetic variants acting in concert to cause disease. They also discovered that gene variants previously identified as causing mitochondrial dysfunction may actually be benign. They detected 44 such variants among their sample population, yet only six of those explained the patients' symptoms.
Read the study:
Calvo SE, Compton AG, Hershman SG, Lim SC, Lieber DS, Tucker EJ, Laskowski A, Garone C, Liu S, Jaffe DB, Christodoulou J, Fletcher JM, Bruno DL, Goldblatt J, Dimauro S, Thorburn DR, Mootha VK. Molecular diagnosis of infantile mitochondrial disease with targeted next-generation sequencing. Sci Transl Med, 4(118):118ra10. 2012. [PubMed]
