Epigenomics Fact Sheet
Epigenomics is a field in which researchers chart the locations and understand the functions of all the chemical tags that mark the genome.
What is the epigenome?
The epigenome is a multitude of chemical compounds that can tell the genome what to do. The human genome is the complete assembly of DNA (deoxyribonucleic acid)-about 3 billion base pairs - that makes each individual unique. DNA holds the instructions for building the proteins that carry out a variety of functions in a cell. The epigenome is made up of chemical compounds and proteins that can attach to DNA and direct such actions as turning genes on or off, controlling the production of proteins in particular cells.When epigenomic compounds attach to DNA and modify its function, they are said to have "marked" the genome. These marks do not change the sequence of the DNA. Rather, they change the way cells use the DNA's instructions. The marks are sometimes passed on from cell to cell as cells divide. They also can be passed down from one generation to the next.
-
What is the epigenome?
The epigenome is a multitude of chemical compounds that can tell the genome what to do. The human genome is the complete assembly of DNA (deoxyribonucleic acid)-about 3 billion base pairs - that makes each individual unique. DNA holds the instructions for building the proteins that carry out a variety of functions in a cell. The epigenome is made up of chemical compounds and proteins that can attach to DNA and direct such actions as turning genes on or off, controlling the production of proteins in particular cells.When epigenomic compounds attach to DNA and modify its function, they are said to have "marked" the genome. These marks do not change the sequence of the DNA. Rather, they change the way cells use the DNA's instructions. The marks are sometimes passed on from cell to cell as cells divide. They also can be passed down from one generation to the next.
What does the epigenome do?
A human being has trillions of cells, specialized for different functions in muscles, bones and the brain, and each of these cells carries essentially the same genome in its nucleus. The differences among cells are determined by how and when different sets of genes are turned on or off in various kinds of cells. Specialized cells in the eye turn on genes that make proteins that can detect light, while specialized cells in red blood cells make proteins that carry oxygen from the air to the rest of the body. The epigenome controls many of these changes to the genome.
-
What does the epigenome do?
A human being has trillions of cells, specialized for different functions in muscles, bones and the brain, and each of these cells carries essentially the same genome in its nucleus. The differences among cells are determined by how and when different sets of genes are turned on or off in various kinds of cells. Specialized cells in the eye turn on genes that make proteins that can detect light, while specialized cells in red blood cells make proteins that carry oxygen from the air to the rest of the body. The epigenome controls many of these changes to the genome.

What makes up the epigenome?
The epigenome is the set of chemical modifications to the DNA and DNA-associated proteins in the cell, which alter gene expression, and are heritable (via meiosis and mitosis). The modifications occur as a natural process of development and tissue differentiation, and can be altered in response to environmental exposures or disease.
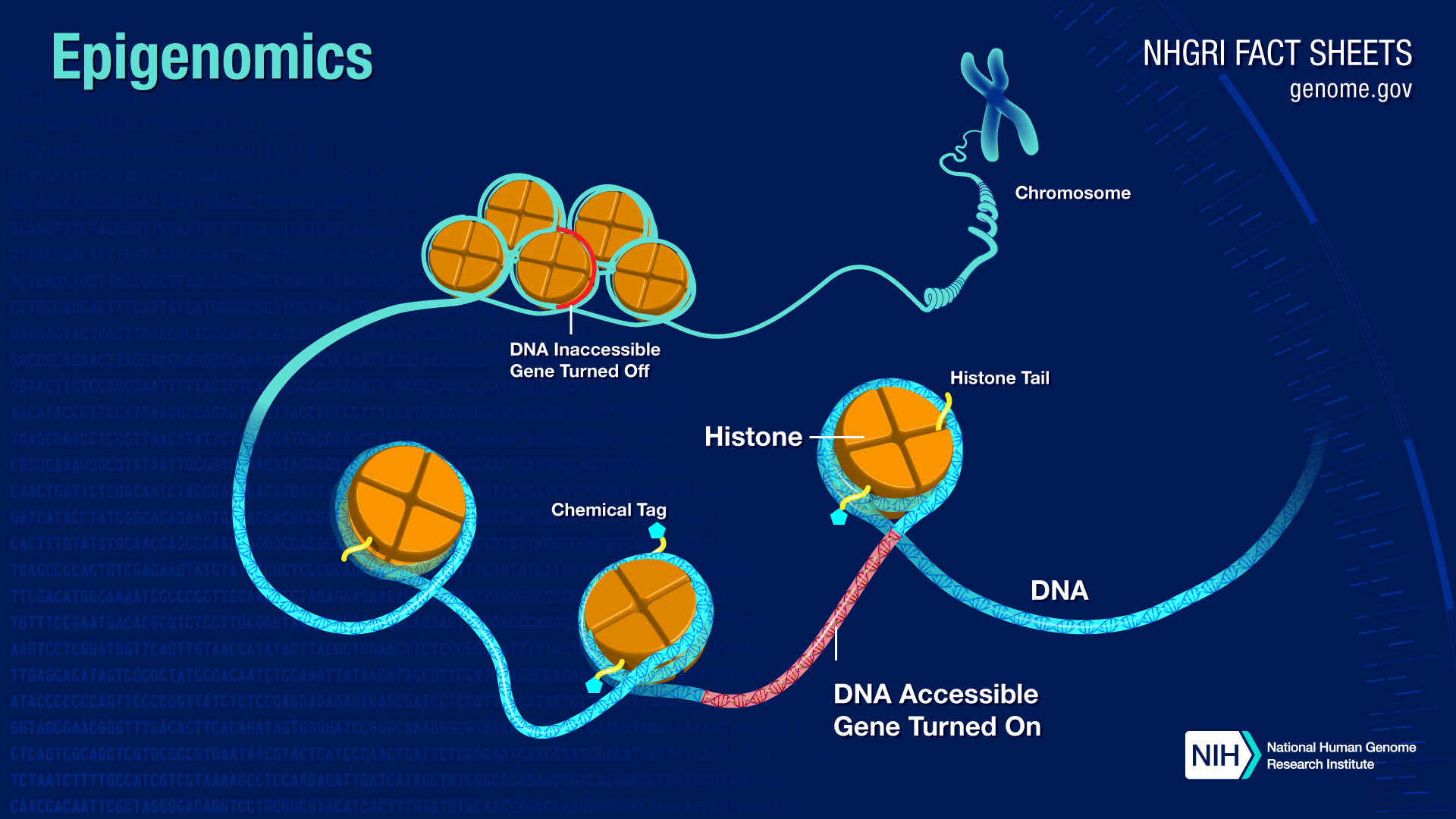
The first type of mark, called DNA methylation, directly affects the DNA in a genome. In this process, proteins attach chemical tags called methyl groups to the bases of the DNA molecule in specific places. The methyl groups turn genes on or off by affecting interactions between the DNA and other proteins. In this way, cells can remember which genes are on or off.The second kind of mark, called histone modification, affects DNA indirectly. DNA in cells is wrapped around histone proteins, which form spool-like structures that enable DNA's very long molecules to be wound up neatly into chromosomes inside the cell nucleus. Proteins can attach a variety of chemical tags to histones. Other proteins in cells can detect these tags and determine whether that region of DNA should be used or ignored in that cell.
-
What makes up the epigenome?
The epigenome is the set of chemical modifications to the DNA and DNA-associated proteins in the cell, which alter gene expression, and are heritable (via meiosis and mitosis). The modifications occur as a natural process of development and tissue differentiation, and can be altered in response to environmental exposures or disease.
The first type of mark, called DNA methylation, directly affects the DNA in a genome. In this process, proteins attach chemical tags called methyl groups to the bases of the DNA molecule in specific places. The methyl groups turn genes on or off by affecting interactions between the DNA and other proteins. In this way, cells can remember which genes are on or off.The second kind of mark, called histone modification, affects DNA indirectly. DNA in cells is wrapped around histone proteins, which form spool-like structures that enable DNA's very long molecules to be wound up neatly into chromosomes inside the cell nucleus. Proteins can attach a variety of chemical tags to histones. Other proteins in cells can detect these tags and determine whether that region of DNA should be used or ignored in that cell.
Is the epigenome inherited?
The genome is passed from parents to their offspring and from cells, when they divide, to their next generation. Much of the epigenome is reset when parents pass their genomes to their offspring; however, under some circumstances, some of the chemical tags on the DNA and histones of eggs and sperm may be passed on to the next generation. When cells divide, often much of the epigenome is passed on to the next generation of cells, helping the cells remain specialized.
-
Is the epigenome inherited?
The genome is passed from parents to their offspring and from cells, when they divide, to their next generation. Much of the epigenome is reset when parents pass their genomes to their offspring; however, under some circumstances, some of the chemical tags on the DNA and histones of eggs and sperm may be passed on to the next generation. When cells divide, often much of the epigenome is passed on to the next generation of cells, helping the cells remain specialized.
What is imprinting?
The human genome contains two copies of every gene-one copy inherited from the mother and one from the father. For a small number of genes, only the copy from the mother gets switched on; for others, only the copy from the father is turned on. This pattern is called imprinting. The epigenome distinguishes between the two copies of an imprinted gene and determines which is switched on.Some diseases are caused by abnormal imprinting. They include Beckwith-Wiedemann syndrome, a disorder associated with body overgrowth and increased risk of cancer; Prader-Willi syndrome, associated with poor muscle tone and constant hunger, leading to obesity; and Angelman syndrome, which leads to intellectual disability, as well motion difficulties.
-
What is imprinting?
The human genome contains two copies of every gene-one copy inherited from the mother and one from the father. For a small number of genes, only the copy from the mother gets switched on; for others, only the copy from the father is turned on. This pattern is called imprinting. The epigenome distinguishes between the two copies of an imprinted gene and determines which is switched on.Some diseases are caused by abnormal imprinting. They include Beckwith-Wiedemann syndrome, a disorder associated with body overgrowth and increased risk of cancer; Prader-Willi syndrome, associated with poor muscle tone and constant hunger, leading to obesity; and Angelman syndrome, which leads to intellectual disability, as well motion difficulties.
Can the epigenome change?
Although all cells in the body contain essentially the same genome, the DNA marked by chemical tags on the DNA and histones gets rearranged when cells become specialized. The epigenome can also change throughout a person's lifetime.
-
Can the epigenome change?
Although all cells in the body contain essentially the same genome, the DNA marked by chemical tags on the DNA and histones gets rearranged when cells become specialized. The epigenome can also change throughout a person's lifetime.
What makes the epigenome change?
Lifestyle and environmental factors (such as smoking, diet and infectious disease) can expose a person to pressures that prompt chemical responses. These responses, in turn, often lead to changes in the epigenome, some of which can be damaging. However, the ability of the epigenome to adjust to the pressures of life appears to be required for normal human health. Some human diseases are caused by malfunctions in the proteins that "read" and "write" epigenomic marks.
-
What makes the epigenome change?
Lifestyle and environmental factors (such as smoking, diet and infectious disease) can expose a person to pressures that prompt chemical responses. These responses, in turn, often lead to changes in the epigenome, some of which can be damaging. However, the ability of the epigenome to adjust to the pressures of life appears to be required for normal human health. Some human diseases are caused by malfunctions in the proteins that "read" and "write" epigenomic marks.
How do changes in the epigenome contribute to cancer?
Cancers are caused by changes in the genome, the epigenome, or both. Changes in the epigenome can switch on or off genes involved in cell growth or the immune response. These changes can lead to uncontrolled growth, a hallmark of cancer, or to a failure of the immune system to destroy tumors.
In a type of brain tumor called glioblastoma, doctors have had some success in treating patients with the drug temozolomide, which kills cancer cells by adding methyl groups to DNA. In some cases, methylation has a welcome secondary effect: it blocks a gene that counteracts temozolomide. Glioblastoma patients whose tumors have such methylated genes are far more likely to respond to temozolomide than those with unmethylated genes.
Changes in the epigenome also can activate growth-promoting genes in stomach cancer, colon cancer and the most common type of kidney cancer. In some other cancers, changes in the epigenome silence genes that normally serve to keep cell growth in check.
To compile a complete list of possible epigenomic changes that can lead to cancer, researchers in The Cancer Genome Atlas (TCGA) Network, which is supported by the National Institutes of Health (NIH), are comparing the genomes and epigenomes of normal cells with those of cancer cells. Among other things, they are looking for changes in the DNA sequence and changes in the number of methyl groups on the DNA.Understanding all the changes that turn a normal cell into a cancer cell will speed efforts to develop new and better ways of diagnosing, treating and preventing cancer.
-
How do changes in the epigenome contribute to cancer?
Cancers are caused by changes in the genome, the epigenome, or both. Changes in the epigenome can switch on or off genes involved in cell growth or the immune response. These changes can lead to uncontrolled growth, a hallmark of cancer, or to a failure of the immune system to destroy tumors.
In a type of brain tumor called glioblastoma, doctors have had some success in treating patients with the drug temozolomide, which kills cancer cells by adding methyl groups to DNA. In some cases, methylation has a welcome secondary effect: it blocks a gene that counteracts temozolomide. Glioblastoma patients whose tumors have such methylated genes are far more likely to respond to temozolomide than those with unmethylated genes.
Changes in the epigenome also can activate growth-promoting genes in stomach cancer, colon cancer and the most common type of kidney cancer. In some other cancers, changes in the epigenome silence genes that normally serve to keep cell growth in check.
To compile a complete list of possible epigenomic changes that can lead to cancer, researchers in The Cancer Genome Atlas (TCGA) Network, which is supported by the National Institutes of Health (NIH), are comparing the genomes and epigenomes of normal cells with those of cancer cells. Among other things, they are looking for changes in the DNA sequence and changes in the number of methyl groups on the DNA.Understanding all the changes that turn a normal cell into a cancer cell will speed efforts to develop new and better ways of diagnosing, treating and preventing cancer.
How are researchers exploring the epigenome?
In a field of study known as epigenomics, researchers are trying to chart the locations and understand the functions of all the chemical tags that mark the genome.
Until recently, scientists thought that human diseases were caused mainly by changes in DNA sequence, infectious agents such as bacteria and viruses, or environmental agents. Now, however, researchers have demonstrated that changes in the epigenome also can cause, or result from, disease. Epigenomics, thus, has become a vital part of efforts to better understand the human body and to improve human health. Epigenomic maps may someday enable doctors to determine an individual's health status and tailor a patient's response to therapies.
As part of the ENCODE (ENCyclopedia Of DNA Elements) project-which aims to catalog the working parts of the genome-the National Human Genome Research Institute is funding researchers to make epigenomic maps of various cell types. Other NIH-supported investigators have developed a number of epigenomic maps from several human organs and tissues. These NIH projects are part of an international effort to understand how epigenomics could lead to better prevention, diagnosis and treatment of disease.
-
How are researchers exploring the epigenome?
In a field of study known as epigenomics, researchers are trying to chart the locations and understand the functions of all the chemical tags that mark the genome.
Until recently, scientists thought that human diseases were caused mainly by changes in DNA sequence, infectious agents such as bacteria and viruses, or environmental agents. Now, however, researchers have demonstrated that changes in the epigenome also can cause, or result from, disease. Epigenomics, thus, has become a vital part of efforts to better understand the human body and to improve human health. Epigenomic maps may someday enable doctors to determine an individual's health status and tailor a patient's response to therapies.
As part of the ENCODE (ENCyclopedia Of DNA Elements) project-which aims to catalog the working parts of the genome-the National Human Genome Research Institute is funding researchers to make epigenomic maps of various cell types. Other NIH-supported investigators have developed a number of epigenomic maps from several human organs and tissues. These NIH projects are part of an international effort to understand how epigenomics could lead to better prevention, diagnosis and treatment of disease.
Last updated: August 16, 2020
