Cost Categories
The expenditures included in each category were established based on discussions between NHGRI staff and sequencing center personnel.
For the two graphs ("Cost per Megabase of DNA Sequence" and "Cost per Genome"), the following 'production' costs are accounted for:
- Labor, administration, management, utilities, reagents, and consumables
- Sequencing instruments and other large equipment (amortized over three years)
- Informatics activities directly related to sequence production (e.g., laboratory information management systems and initial data processing)
- Submission of data to a public database
- Indirect Costs as they relate to the above items
In the case of costs covered by significant subsidies to a sequencing center (e.g., a grantee institution providing funds for purchasing large equipment), NHGRI has attempted to appropriately account for such costs in these analyses.
The costs associated with the following 'non-production' activities are not reflected in the two graphs:
- Quality assessment/control for sequencing projects
- Technology development to improve sequencing pipelines
- Development of bioinformatics/computational tools to improve sequencing pipelines or to improve downstream sequence analysis
- Management of individual sequencing projects
- Informatics equipment
- Data analysis downstream of initial data processing (e.g., sequence assembly, sequence alignments, identifying variants, and interpretation of results)
DNA Sequencing Technologies
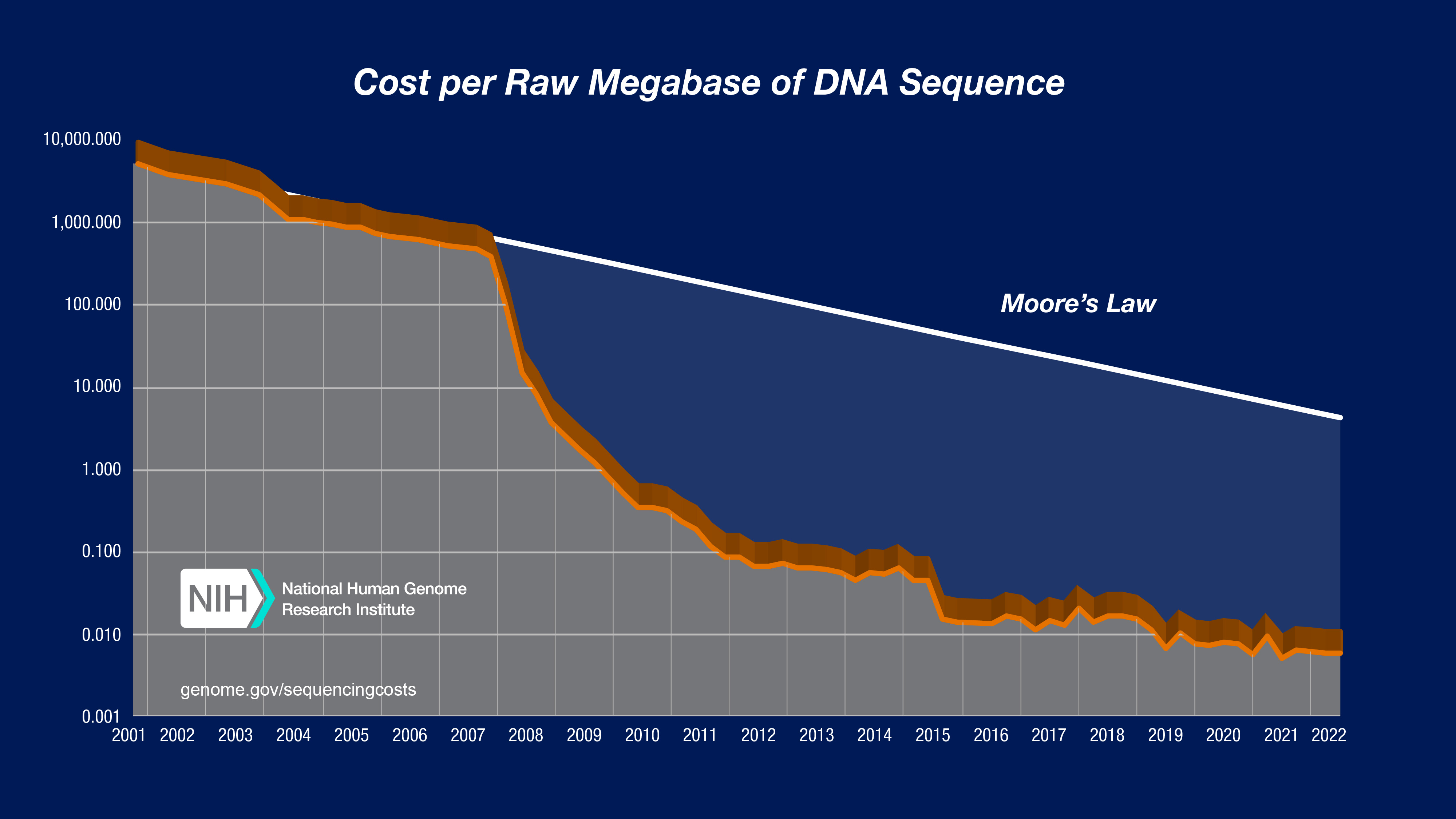
In both graphs, the data from 2001 through October 2007 represent the costs of generating DNA sequence using Sanger-based chemistries and capillary-based instruments ('first generation' sequencing platforms). Beginning in January 2008, the data represent the costs of generating DNA sequence using 'second-generation' (or 'next-generation') sequencing platforms. The change in instruments represents the rapid evolution of DNA sequencing technologies that has occurred in recent years.
Quality
For the Sanger-based sequence data, the cost accounting reflects the generation of bases with a minimum quality score of Phred20(or Q20), which represents an error probability of 1 % and is an accepted community standard for a high-quality base. For sequence data generated with second-generation sequencing platforms, there is not yet a single accepted measure of accuracy; each manufacturer provides quality scores that are, at this time, accepted by the NHGRI sequencing centers as equivalent to or greater than Q20.
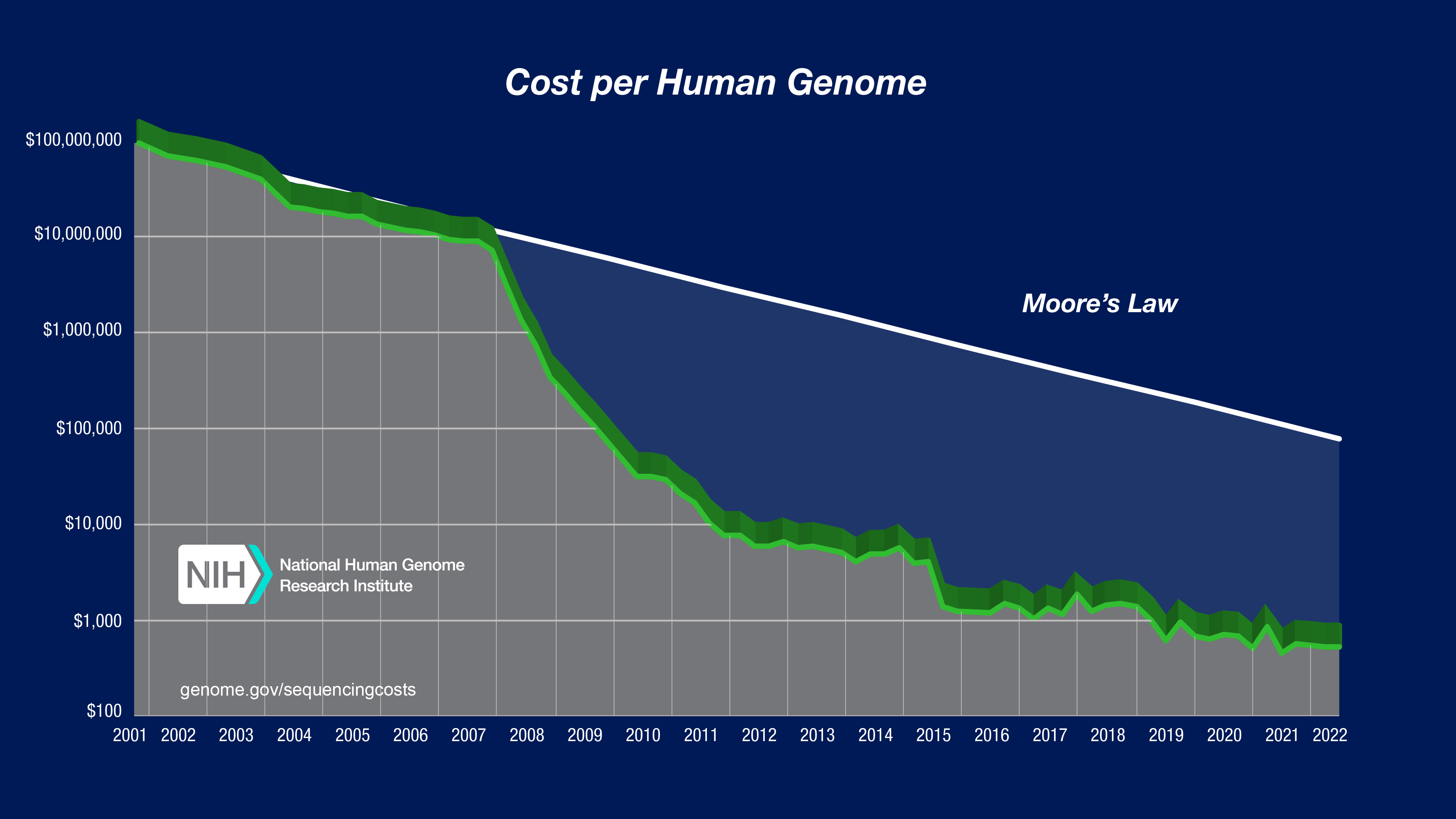
In the "Cost per Megabase of DNA Sequence" graph, the data reflect the cost of generating raw, unassembled sequence data; no adjustment was made for data generated using different instruments despite significant differences in the sequence read lengths. In contrast, the "Cost per Genome" graph does take these differences into account since sequence read length influences the ability to generate an assembled genome sequence.
Genome Coverage
The "Cost per Genome" graph was generated using the same underlying data as that used to generate the "Cost per Megabase of DNA Sequence" graph; the former thus reflects an estimate of the cost of sequencing a human-sized genome rather than the actual costs for specific genome-sequencing projects.
To calculate the cost for sequencing a genome, one needs to know the size of that genome and the required 'sequence coverage' (i.e., 'sequence redundancy') to generate a high-quality assembly of the genome given the specific sequencing platform being used. For generating the "Cost per Genome" graph, the assumed genome size was 3,000 Mb (i.e., the size of a human genome). The assumed sequence coverage needed differed among sequencing platforms, depending on the average sequence read length for that platform.
The following 'sequence coverage' values were used in calculating the cost per genome:
- Sanger-based sequencing (average read length=500-600 bases): 6-fold coverage
- 454 sequencing (average read length=300-400 bases): 10-fold coverage
- Illumina and SOLiD sequencing (average read length=75-150 bases): 30-fold coverage
For data since January 2008 (representing data generated using 'second-generation' sequencing platforms), the "Cost per Genome" graph reflects projects involving the 're-sequencing' of the human genome, where an available reference human genome sequence is available to serve as a backbone for downstream data analyses. The required 'sequence coverage' would be greater for sequencing genomes for which no reference genome sequence is available.