Last updated: March 18, 2013
Bioinformatics: Introduction
Bioinformatics: Introduction
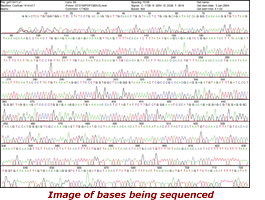 When the Human Genome Project was begun in 1990 it was understood that to meet the project's goals, the speed of DNA sequencing would have to increase and the cost would have to come down. Over the life of the project virtually every aspect of DNA sequencing was improved. It took the project approximately four years to sequence its first one billion bases but just four months to sequence the second billion bases.
When the Human Genome Project was begun in 1990 it was understood that to meet the project's goals, the speed of DNA sequencing would have to increase and the cost would have to come down. Over the life of the project virtually every aspect of DNA sequencing was improved. It took the project approximately four years to sequence its first one billion bases but just four months to sequence the second billion bases.
During the month of January, 2003, 1.5 billion bases were sequenced. As the speed of DNA sequencing increased, the cost decreased from 10 dollars per base in 1990 to 10 cents per base at the conclusion of the project in April 2003. Although the Human Genome Project is officially over, improvements in DNA sequencing continue to be made. Researchers are experimenting with new methods for sequencing DNA that have the potential to sequence a human genome in just a matter of weeks for a few thousand dollars.
 DNA sequencing performed on an industrial scale has produced a vast amount of data to analyze. In August 2005 it was announced that the three largest public collections of DNA and RNA sequences together store one hundred billion bases, representing over 165,000 different organisms. As sequence data began to pile up, the need for new and better methods of sequence analysis was critical.
DNA sequencing performed on an industrial scale has produced a vast amount of data to analyze. In August 2005 it was announced that the three largest public collections of DNA and RNA sequences together store one hundred billion bases, representing over 165,000 different organisms. As sequence data began to pile up, the need for new and better methods of sequence analysis was critical.
Bioinformatics is the branch of biology that is concerned with the acquisition, storage, and analysis of the information found in nucleic acid and protein sequence data. Computers and bioinformatics software are the tools of the trade.
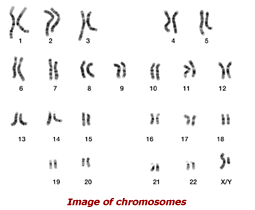 Genetic data represent a treasure trove for researchers and companies interested in how genes contribute to our health and well being. Almost half of the genes identified by the Human Genome Project have no known function. Researchers are using bioinformatics to identify genes, establish their functions, and develop gene-based strategies for preventing, diagnosing, and treating disease.
Genetic data represent a treasure trove for researchers and companies interested in how genes contribute to our health and well being. Almost half of the genes identified by the Human Genome Project have no known function. Researchers are using bioinformatics to identify genes, establish their functions, and develop gene-based strategies for preventing, diagnosing, and treating disease.
 A DNA sequencing reaction produces a sequence that is several hundred bases long. Gene sequences typically run for thousands of bases. The largest known gene is that associated with Duchenne muscular dystrophy. It is approximately 2.4 million bases in length. In order to study genes, scientists first assemble long DNA sequences from series of shorter overlapping sequences.
A DNA sequencing reaction produces a sequence that is several hundred bases long. Gene sequences typically run for thousands of bases. The largest known gene is that associated with Duchenne muscular dystrophy. It is approximately 2.4 million bases in length. In order to study genes, scientists first assemble long DNA sequences from series of shorter overlapping sequences.
Scientists enter their assembled sequences into genetic databases so that other scientists may use the data. Since the sequences of the two DNA strands are complementary, it is only necessary to enter the sequence of one DNA strand into a database. By selecting an appropriate computer program, scientists can use sequence data to look for genes, get clues to gene functions, examine genetic variation, and explore evolutionary relationships. Bioinformatics is a young and dynamic science. New bioinformatic software is being developed while existing software is continually updated.
