Last updated: March 05, 2015
Bioinformatics: Finding Genes
Bioinformatics: Finding Genes
Click on the figures below to view enlarged images.
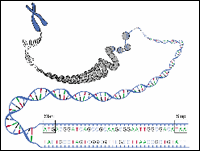
Figure 1: DNA Sequences- three bases and stop codons
One of the most important aspects of bioinformatics is identifying genes within a long DNA sequence. Until the development of bioinformatics, the only way to locate genes along the chromosome was to study their behavior in the organism (in vivo) or isolate the DNA and study it in a test tube (in vitro). Bioinformatics allows scientists to make educated guesses about where genes are located simply by analyzing sequence data using a computer (in silico).
In principle, locating genes should be easy. DNA sequences that code for proteins begin with the three bases ATG that code for the amino acid methionine and they end with one or more stop codons; either TAA, TAG or TGA. Unfortunately, finding genes isn't always so easy.
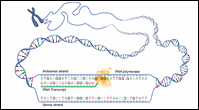
Figure 2: Sense Strand / Antisense Strand
Let's consider a DNA sequence that contains a gene of interest. The DNA strand that codes for the protein is called the sense strand because its sequence reads the same as that of the messenger RNA. The other strand is called the antisense strand and serves as the template for RNA polymerase during transcription.
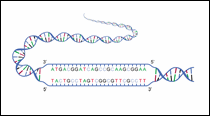
Figure 3: Open Reading Frame
A gene begins with a codon for the amino acid methionine and ends with one of three stop codons. The codons between the start and stop signals code for the various amino acids of the gene product but do not include any of the three stop codons. When examining an unknown DNA sequence, one indication that it may be part of a gene is the presence of an open reading frame or ORF. An ORF is any stretch of DNA that when transcribed into RNA has no stop codon.
Figure 4: Three Different Reading Frames
A computer program can be used to check an unknown DNA sequence for ORFs. The program transcribes each DNA strand into its complementary RNA sequence and then translates the RNA sequence into an amino acid sequence. Each DNA strand can be read in three different reading frames. This means that the computer must perform six different translations for any given double-stranded DNA sequence.
Figure 5: Regions of DNA sequence that might be part of genes
The presence of an ORF doesn't guarantee that the DNA sequence is part of a gene. We expect that, just by chance, there will be some long stretches of DNA that do not contain stop codons yet are not parts of genes. Likewise, codons for methionine do not always mark the start of a gene sequence. Methionine codons are also found within genes. Nevertheless, searching for ORFs identifies regions of the DNA sequence that might be parts of genes.
Figure 6: Strands with 5' and 3'
A single RNA or DNA strand has a phosphate group at one end and a sugar (ribose for RNA and deoxyribose for DNA) at the other end. The end of the strand with the phosphate group is called the 5' end and the opposite end with the sugar is called the 3' end. In the double helix, the two strands run in opposite directions. That is, one strand runs in the 5' to 3' direction while the complementary strand runs in the 3' to 5' direction.
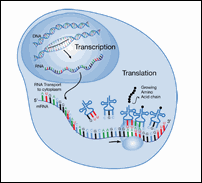
Figure 7: Transcription and Translation
The enzymes and ribosomes that carry out protein synthesis only work in one direction. During transcription, the mRNA is made in the 5' to 3' direction. During translation, the mRNA is read in the 5' to 3' direction. This means that a computer program looking for ORFs also must read each DNA strand in the 5' to 3' direction.
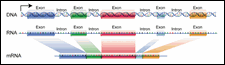
Figure 8: Exons and Introns
It is easier to locate genes in bacterial DNA than in eukaryotic DNA. In bacteria, the genes are arranged like beads on a string. Each gene consists of a single ORF. The situation in eukaryotic organisms is complicated by the split nature of the genes. Most eukaryotic genes take the form of alternating exons and introns. Each exon is an ORF that codes for amino acids. The intron sequences do not code for amino acids and contain internal stop codons.
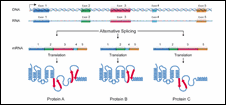
Figure 9: Alternative Splicing
One of the surprises of the Human Genome Project was the relatively small number of genes found - about 25,000. One might ask, "How can something as complicated as a human have only 25 percent more genes than the tiny roundworm C. elegans?" Part of the answer seems to involve alternative splicing. Alternative splicing refers to the process by which a given gene is spliced into more than one type of mRNA molecule.
ORFs are just one feature that a computer program looks for when locating potential genes. Genes are also characterized by specific control sequences that are recognized by enzymes involved with transcription and translation. When a computer program finds a DNA sequence that satisfies all of these gene features (an ORF plus the appropriate control sequences), it identifies the sequence as likely coming from a gene. Only testing the DNA sequence in the laboratory can prove that the gene is active in an organism however.