DNA Sequencing Fact Sheet
DNA sequencing determines the order of the four chemical building blocks - called "bases" - that make up the DNA molecule.
What is DNA sequencing?
Sequencing DNA means determining the order of the four chemical building blocks - called "bases" - that make up the DNA molecule. The sequence tells scientists the kind of genetic information that is carried in a particular DNA segment. For example, scientists can use sequence information to determine which stretches of DNA contain genes and which stretches carry regulatory instructions, turning genes on or off. In addition, and importantly, sequence data can highlight changes in a gene that may cause disease.
In the DNA double helix, the four chemical bases always bond with the same partner to form "base pairs." Adenine (A) always pairs with thymine (T); cytosine (C) always pairs with guanine (G). This pairing is the basis for the mechanism by which DNA molecules are copied when cells divide, and the pairing also underlies the methods by which most DNA sequencing experiments are done. The human genome contains about 3 billion base pairs that spell out the instructions for making and maintaining a human being.
-
What is DNA sequencing?
Sequencing DNA means determining the order of the four chemical building blocks - called "bases" - that make up the DNA molecule. The sequence tells scientists the kind of genetic information that is carried in a particular DNA segment. For example, scientists can use sequence information to determine which stretches of DNA contain genes and which stretches carry regulatory instructions, turning genes on or off. In addition, and importantly, sequence data can highlight changes in a gene that may cause disease.
In the DNA double helix, the four chemical bases always bond with the same partner to form "base pairs." Adenine (A) always pairs with thymine (T); cytosine (C) always pairs with guanine (G). This pairing is the basis for the mechanism by which DNA molecules are copied when cells divide, and the pairing also underlies the methods by which most DNA sequencing experiments are done. The human genome contains about 3 billion base pairs that spell out the instructions for making and maintaining a human being.
How new is DNA sequencing?
Since the completion of the Human Genome Project, technological improvements and automation have increased speed and lowered costs to the point where individual genes can be sequenced routinely, and some labs can sequence well over 100,000 billion bases per year, and an entire genome can be sequenced for just a few thousand dollars.
Many of these new technologies were developed with support from the National Human Genome Research Institute (NHGRI) Genome Technology Program and its Advanced DNA Sequencing Technology awards. One of NHGRI's goals is to promote new technologies that could eventually reduce the cost of sequencing a human genome of even higher quality than is possible today and for less than $1,000.
-
How new is DNA sequencing?
Since the completion of the Human Genome Project, technological improvements and automation have increased speed and lowered costs to the point where individual genes can be sequenced routinely, and some labs can sequence well over 100,000 billion bases per year, and an entire genome can be sequenced for just a few thousand dollars.
Many of these new technologies were developed with support from the National Human Genome Research Institute (NHGRI) Genome Technology Program and its Advanced DNA Sequencing Technology awards. One of NHGRI's goals is to promote new technologies that could eventually reduce the cost of sequencing a human genome of even higher quality than is possible today and for less than $1,000.

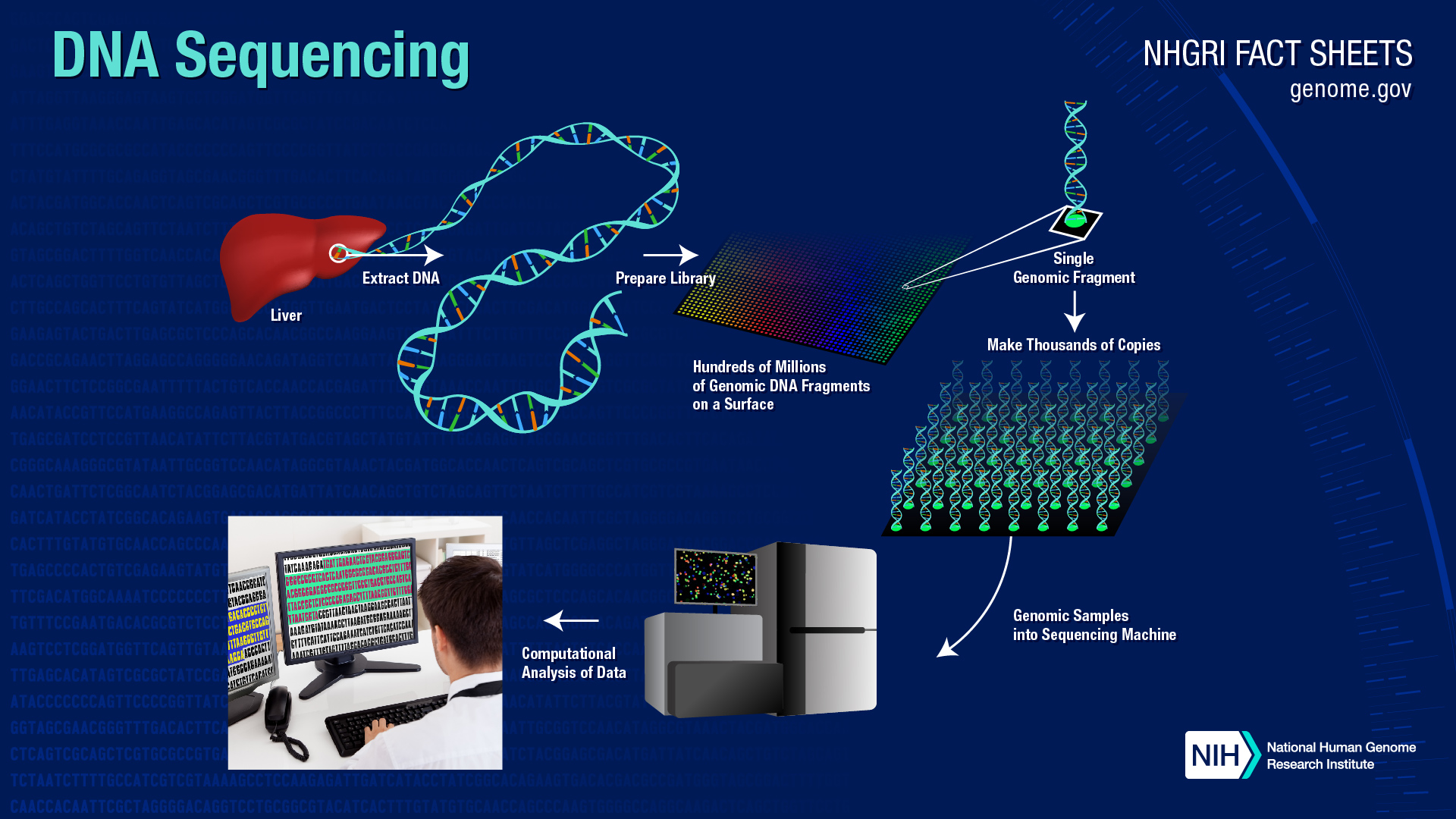
What new sequencing methods have been developed?
Since the completion of the Human Genome Project, technological improvements and automation have increased speed and lowered costs to the point where individual genes can be sequenced routinely, and some labs can sequence well over 100,000 billion bases per year, and an entire genome can be sequenced for just a few thousand dollars.Many of these new technologies were developed with support from the National Human Genome Research Institute (NHGRI) Genome Technology Program and its Advanced DNA Sequencing Technology awards. One of NHGRI's goals is to promote new technologies that could eventually reduce the cost of sequencing a human genome of even higher quality than is possible today and for less than $1,000.
-
What new sequencing methods have been developed?
Since the completion of the Human Genome Project, technological improvements and automation have increased speed and lowered costs to the point where individual genes can be sequenced routinely, and some labs can sequence well over 100,000 billion bases per year, and an entire genome can be sequenced for just a few thousand dollars.Many of these new technologies were developed with support from the National Human Genome Research Institute (NHGRI) Genome Technology Program and its Advanced DNA Sequencing Technology awards. One of NHGRI's goals is to promote new technologies that could eventually reduce the cost of sequencing a human genome of even higher quality than is possible today and for less than $1,000.
Are newer sequencing technologies under development?
One new sequencing technology involves watching DNA polymerase molecules as they copy DNA - the same molecules that make new copies of DNA in our cells - with a very fast movie camera and microscope, and incorporating different colors of bright dyes, one each for the letters A, T, C and G. This method provides different and very valuable information than what's provided by the instrument systems that are in most common use.
Another new technology in development entails the use of nanopores to sequence DNA. Nanopore-based DNA sequencing involves threading single DNA strands through extremely tiny pores in a membrane. DNA bases are read one at a time as they squeeze through the nanopore. The bases are identified by measuring differences in their effect on ions and electrical current flowing through the pore.Using nanopores to sequence DNA offers many potential advantages over current methods. The goal is for sequencing to cost less and be done faster. Unlike sequencing methods currently in use, nanopore DNA sequencing means researchers can study the same molecule over and over again.
-
Are newer sequencing technologies under development?
One new sequencing technology involves watching DNA polymerase molecules as they copy DNA - the same molecules that make new copies of DNA in our cells - with a very fast movie camera and microscope, and incorporating different colors of bright dyes, one each for the letters A, T, C and G. This method provides different and very valuable information than what's provided by the instrument systems that are in most common use.
Another new technology in development entails the use of nanopores to sequence DNA. Nanopore-based DNA sequencing involves threading single DNA strands through extremely tiny pores in a membrane. DNA bases are read one at a time as they squeeze through the nanopore. The bases are identified by measuring differences in their effect on ions and electrical current flowing through the pore.Using nanopores to sequence DNA offers many potential advantages over current methods. The goal is for sequencing to cost less and be done faster. Unlike sequencing methods currently in use, nanopore DNA sequencing means researchers can study the same molecule over and over again.
What do improvements in DNA sequencing mean for human health?
Researchers now are able to compare large stretches of DNA - 1 million bases or more - from different individuals quickly and cheaply. Such comparisons can yield an enormous amount of information about the role of inheritance in susceptibility to disease and in response to environmental influences. In addition, the ability to sequence the genome more rapidly and cost-effectively creates vast potential for diagnostics and therapies.
Although routine DNA sequencing in the doctor's office is still many years away, some large medical centers have begun to use sequencing to detect and treat some diseases. In cancer, for example, physicians are increasingly able to use sequence data to identify the particular type of cancer a patient has. This enables the physician to make better choices for treatments.
Researchers in the NHGRI-supported Undiagnosed Diseases Program use DNA sequencing to try to identify the genetic causes of rare diseases. Other researchers are studying its use in screening newborns for disease and disease risk.
Moreover, The Cancer Genome Atlas project, which is supported by NHGRI and the National Cancer Institute, is using DNA sequencing to unravel the genomic details of some 30 cancer types. Another National Institutes of Health program examines how gene activity is controlled in different tissues and the role of gene regulation in disease. Ongoing and planned large-scale projects use DNA sequencing to examine the development of common and complex diseases, such as heart disease and diabetes, and in inherited diseases that cause physical malformations, developmental delay and metabolic diseases.
Comparing the genome sequences of different types of animals and organisms, such as chimpanzees and yeast, can also provide insights into the biology of development and evolution.
-
What do improvements in DNA sequencing mean for human health?
Researchers now are able to compare large stretches of DNA - 1 million bases or more - from different individuals quickly and cheaply. Such comparisons can yield an enormous amount of information about the role of inheritance in susceptibility to disease and in response to environmental influences. In addition, the ability to sequence the genome more rapidly and cost-effectively creates vast potential for diagnostics and therapies.
Although routine DNA sequencing in the doctor's office is still many years away, some large medical centers have begun to use sequencing to detect and treat some diseases. In cancer, for example, physicians are increasingly able to use sequence data to identify the particular type of cancer a patient has. This enables the physician to make better choices for treatments.
Researchers in the NHGRI-supported Undiagnosed Diseases Program use DNA sequencing to try to identify the genetic causes of rare diseases. Other researchers are studying its use in screening newborns for disease and disease risk.
Moreover, The Cancer Genome Atlas project, which is supported by NHGRI and the National Cancer Institute, is using DNA sequencing to unravel the genomic details of some 30 cancer types. Another National Institutes of Health program examines how gene activity is controlled in different tissues and the role of gene regulation in disease. Ongoing and planned large-scale projects use DNA sequencing to examine the development of common and complex diseases, such as heart disease and diabetes, and in inherited diseases that cause physical malformations, developmental delay and metabolic diseases.
Comparing the genome sequences of different types of animals and organisms, such as chimpanzees and yeast, can also provide insights into the biology of development and evolution.
Last updated: June 27, 2023
