Transcriptome Fact Sheet
A transcriptome is a collection of all the gene readouts present in a cell.
What is a transcriptome?
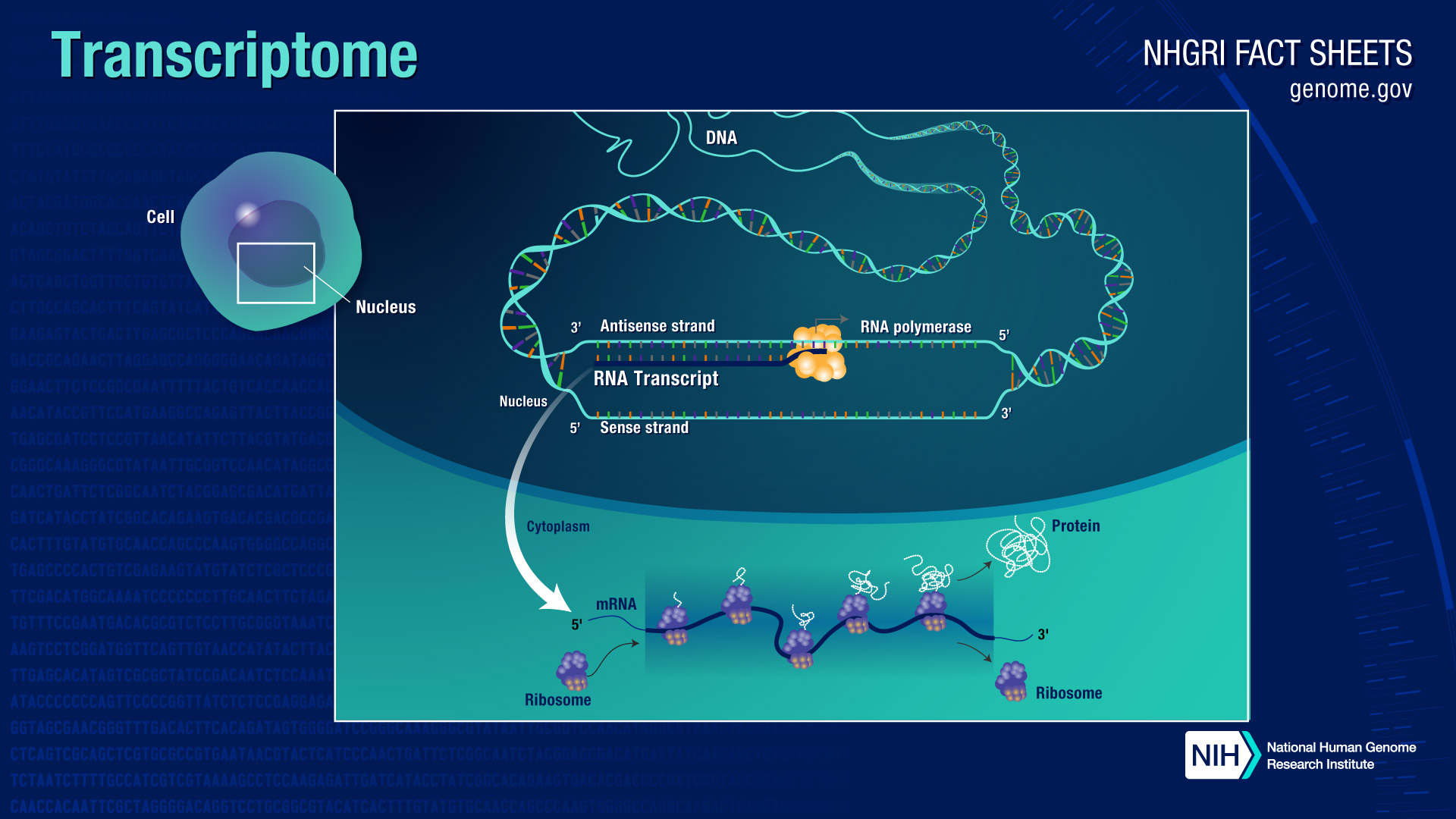
The human genome is made up of DNA (deoxyribonucleic acid), a long, winding molecule that contains the instructions needed to build and maintain cells. These instructions are spelled out in the form of "base pairs" of four different chemicals, organized into 20,000 to 25,000 genes. For the instructions to be carried out, DNA must be "read" and transcribed - in other words, copied - into RNA (ribonucleic acid). These gene readouts are called transcripts, and a transcriptome is a collection of all the gene readouts present in a cell.
There are various kinds of RNA. The major type, called messenger RNA (mRNA), plays a vital role in making proteins. In this process: mRNA is transcribed from genes; then the mRNA transcripts are delivered to ribosomes, the molecular machines located in the cell's cytoplasm; then the ribosomes read, or "translate," the sequence of chemical letters in the mRNA and assemble building blocks called amino acids into proteins.
DNA can also be transcribed into other types of RNA that do not code for proteins. Such transcripts may serve to influence cell structure and to regulate genes.
-
What is a transcriptome?
The human genome is made up of DNA (deoxyribonucleic acid), a long, winding molecule that contains the instructions needed to build and maintain cells. These instructions are spelled out in the form of "base pairs" of four different chemicals, organized into 20,000 to 25,000 genes. For the instructions to be carried out, DNA must be "read" and transcribed - in other words, copied - into RNA (ribonucleic acid). These gene readouts are called transcripts, and a transcriptome is a collection of all the gene readouts present in a cell.
There are various kinds of RNA. The major type, called messenger RNA (mRNA), plays a vital role in making proteins. In this process: mRNA is transcribed from genes; then the mRNA transcripts are delivered to ribosomes, the molecular machines located in the cell's cytoplasm; then the ribosomes read, or "translate," the sequence of chemical letters in the mRNA and assemble building blocks called amino acids into proteins.
DNA can also be transcribed into other types of RNA that do not code for proteins. Such transcripts may serve to influence cell structure and to regulate genes.
What can a transcriptome tell us?
An RNA sequence mirrors the sequence of the DNA from which it was transcribed. Consequently, by analyzing the entire collection of RNA sequences in a cell (the transcriptome) researchers can determine when and where each gene is turned on or off in the cells and tissues of an organism.
Depending on the technique used, it is often possible to count the number of transcripts to determine the amount of gene activity - also called gene expression - in a certain cell or tissue type.
In humans and other organisms, nearly every cell contains the same genes, but different cells show different patterns of gene expression. These differences are responsible for the many different properties and behaviors of various cells and tissues, both in health and disease.
By collecting and comparing transcriptomes of different types of cells, researchers can gain a deeper understanding of what constitutes a specific cell type, how that type of cell normally functions, and how changes in the normal level of gene activity may reflect or contribute to disease. In addition, transcriptomes may enable researchers to generate a comprehensive, genome-wide picture of what genes are active in which cells.
-
What can a transcriptome tell us?
An RNA sequence mirrors the sequence of the DNA from which it was transcribed. Consequently, by analyzing the entire collection of RNA sequences in a cell (the transcriptome) researchers can determine when and where each gene is turned on or off in the cells and tissues of an organism.
Depending on the technique used, it is often possible to count the number of transcripts to determine the amount of gene activity - also called gene expression - in a certain cell or tissue type.
In humans and other organisms, nearly every cell contains the same genes, but different cells show different patterns of gene expression. These differences are responsible for the many different properties and behaviors of various cells and tissues, both in health and disease.
By collecting and comparing transcriptomes of different types of cells, researchers can gain a deeper understanding of what constitutes a specific cell type, how that type of cell normally functions, and how changes in the normal level of gene activity may reflect or contribute to disease. In addition, transcriptomes may enable researchers to generate a comprehensive, genome-wide picture of what genes are active in which cells.

How transcriptomes be used to explore gene function?
The function of most genes is not yet known. A search of a transcriptome database can give researchers a list of all the tissues in which a gene is expressed, providing clues about its possible function.
For example, if the transcriptome database shows that an unknown gene's expression levels are dramatically higher in cancer cells than in healthy cells, the unknown gene may play a role in cell growth. Or if an unknown gene is expressed in fat tissue but not in bone or muscle tissue, the unknown gene may be involved in fat storage or metabolism. In both instances, the transcriptome data give researchers a good place to start searching for a newly found gene's function.
The National Human Genome Research Institute (NHGRI), which is part of the National Institutes of Health (NIH), has participated in two projects that created transcriptome resources for researchers around the world-the Mammalian Gene Collection initiative and the Mouse Transcriptome Project.
The Mammalian Gene Collection initiative built a free, public library of human, mouse, and rat mRNA sequences. The project was led by NHGRI and the National Cancer Institute (NCI), also part of NIH. The mouse and rat are important models with which to study human biology.
The Mouse Transcriptome Project was an NIH-supported initiative that generated a free, public database of gene transcripts for many mouse tissues. These tissue-specific gene expression data, which are mapped to the mouse genome, are available in a searchable format in the Mouse Reference Transcriptome Database.
Several other transcriptome resources exist, including those in NIH programs such as the Genotype-Tissue Expression Project (GTEx) and the Encyclopedia of DNA Elements (ENCODE). GTEx is creating a catalog of human gene expression in a variety of different tissues. ENCODE researchers aim to characterize and understand the working parts of the genome, including the transcriptome. Both Novartis and the European Molecular Biology Laboratory have well-established gene expression databases.
-
How transcriptomes be used to explore gene function?
The function of most genes is not yet known. A search of a transcriptome database can give researchers a list of all the tissues in which a gene is expressed, providing clues about its possible function.
For example, if the transcriptome database shows that an unknown gene's expression levels are dramatically higher in cancer cells than in healthy cells, the unknown gene may play a role in cell growth. Or if an unknown gene is expressed in fat tissue but not in bone or muscle tissue, the unknown gene may be involved in fat storage or metabolism. In both instances, the transcriptome data give researchers a good place to start searching for a newly found gene's function.
The National Human Genome Research Institute (NHGRI), which is part of the National Institutes of Health (NIH), has participated in two projects that created transcriptome resources for researchers around the world-the Mammalian Gene Collection initiative and the Mouse Transcriptome Project.
The Mammalian Gene Collection initiative built a free, public library of human, mouse, and rat mRNA sequences. The project was led by NHGRI and the National Cancer Institute (NCI), also part of NIH. The mouse and rat are important models with which to study human biology.
The Mouse Transcriptome Project was an NIH-supported initiative that generated a free, public database of gene transcripts for many mouse tissues. These tissue-specific gene expression data, which are mapped to the mouse genome, are available in a searchable format in the Mouse Reference Transcriptome Database.
Several other transcriptome resources exist, including those in NIH programs such as the Genotype-Tissue Expression Project (GTEx) and the Encyclopedia of DNA Elements (ENCODE). GTEx is creating a catalog of human gene expression in a variety of different tissues. ENCODE researchers aim to characterize and understand the working parts of the genome, including the transcriptome. Both Novartis and the European Molecular Biology Laboratory have well-established gene expression databases.
Last updated: August 17, 2020
