Last updated: April 12, 2013
2013 Release Nhgri Celebrates 10th Anniversary Of The Human Genome Project

NHGRI celebrates 10th anniversary of the Human Genome Project
Genomics on productive trajectory a decade after completing flagship project
"On the 10-year anniversary of the completion of the Human Genome Project, it is appropriate to celebrate our accomplishments over the past decide and to reflect on the impact of genomics on research, medicine and society," said NHGRI Director Eric D. Green, M.D., Ph.D. "By improving our understanding of basic genome biology, the genomic underpinnings of disease and genomic medicine, the field gets ever closer to its ultimate goal - improving human health through genomics research."
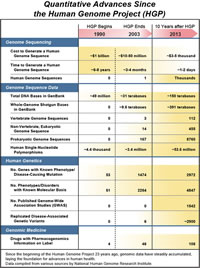
Advances in DNA sequencing technologies have led to plummeting costs for sequencing a human genome, which in turn has yielded major biological insights and new medical applications. Sequencing the first human genome by the HGP took about $1 billion and 13 years, Dr. Green estimates. Today, sequencing a human genome costs less than $5,000 and takes only a day or two.
But sequencing any genome without having the means to interpret it was like looking at Egyptian hieroglyphs before the discovery of the Rosetta Stone. To understand the meaning of the sequence, the field developed approaches for studying genome function, including sequencing the genomes of more than 135 other organisms and conducting the first global surveys of human genomic variation.
Knowledge about the genomes of other organisms has helped researchers read evolution's notebook. By comparing the genome sequences of numerous vertebrates - from chimpanzees to platypuses - and other eukaryotic organisms such as yeast and flat worms, researchers can identify genomic segments with important functions that persisted essentially unchanged over very large evolutionary times. Surprisingly, just 1.5 percent of the human genome actually encodes proteins, but a comparative analysis shows that another 5 to 8 percent, including the genes, is highly conserved over the millennia.
Looking deeper into the genome and its functions, 440 researchers in 32 labs around the world formed the ENCyclopedia Of DNA Elements (ENCODE) consortium. A flurry of ENCODE publications last year revealed new insights about human genome function, including the location of 4 million genomic regions that are candidates for being regulatory switches that turn genes on and off and evidence of biological activity for upwards of 80 percent of the human genome (much of which was once referred to as 'junk DNA').
With the first complete human genome sequence in hand, researchers also set out to measure human genomic variation. Early studies during the HGP suggested that humans differ from each other by only one-tenth of a percent across their 3-billion-base genome. Understanding that variation is critical for establishing the small subset of variants that influence human health and disease.
The International HapMap Project analyzed the genomes of people from Europe, China, Japan and Africa, producing the first catalog of human genomic variation. Biotech companies used the HapMap findings and those from the ongoing 1000 Genomes Project (which aims to extend and refine the HapMap catalog) to build tools for analyzing study populations with and without disease in order to identify variants associated with disease susceptibility. In just a few years, these so-called genome-wide association studies (GWAS) have identified thousands of genomic variations that can affect an individual's likelihood of developing a disease or the underlying pathology of disease progression.
GWAS approaches and other breakthroughs have identified so many clinically relevant genomic variants. Among the utility of such knowledge has been the U.S. Food and Drug Administration requirement to add information on labels of more than 100 existing or new medications about the specific genetic markers. In short, this additional label information now notifies physicians to consider the patient's genomic makeup when prescribing.
The last decade has also revealed the transformative power of using genomic information for the diagnosis and treatment of cancer. For example, the breast cancer drug trastuzumab (Herceptin) only works for women whose tumors have a particular genomic profile, called HER-2 positive. Studies have also found that lung cancer patients whose tumors are positive for EGFR mutations respond to the drugs gefitinib (Iressa) and erlotinib (Tarceva), which target these mutations, but not for lung cancers lacking such mutations. Determining the presence of specific genomic variants also avoids the implementation of ineffective treatments; for example, colon cancer patients whose tumors have a mutation in a gene called KRAS derive little benefit from the drugs cetuximab (Erbitux) and panitumumab (Vectibix).
Using the foundational knowledge provided by the HGP, the discovery of gene alterations causing specific diseases has increased dramatically. In 1990, when the HGP began, mutations in only 53 genes had been shown to cause disease; today, that number is over 2,900. Researchers use such molecular discoveries to develop new diagnostics and treatments. These accomplishments focused on rare diseases, defined as those affecting fewer than 200,000 people, but which cumulatively afflict more than 25 million Americans.
While most rare disorders afflict relatively few individuals, the biological pathways identified through the study of these diseases often provide insights into complex diseases that involve multiple genes. For example, today's widespread use of cholesterol-lowering medications began as a pre-HGP investigation into a rare, hereditary condition called familial hypercholesterolemia.
With the goal of improving health outcomes in mind, researchers continue to develop new tools for advancing genomic research and medical applications of genomics. Initiatives such as NHGRI's Clinically Relevant Variants Resource aim to develop approaches for identifying genomic variants that are clinically relevant for a wide range of disorders; likewise, NHGRI's new Clinical Sequencing Exploratory Research program focuses on the technical, ethical, psychosocial and clinical implications of returning patients' genomic information to healthcare providers and, in turn, their patients.
Researchers have long recognized that the integration of genomics into research and medicine will inevitably affect individuals, families, communities and society. Therefore, an integral part of the Human Genome Project and genomics over the past decade has been the study of the ethical, legal and social implications (ELSI) of this research. NHGRI's ELSI Program represents the largest block of bioethics research funding in the world. In its 23 years, NHGRI's ELSI Program has provided some $300 million to support nearly 500 research projects, which in turn has created a body of knowledge and fostered a community of researchers focused on understanding the risks and benefits of genomic science. Among other issues, the ELSI research community is now focused on the important ethical issues related to the translation of genomic information to the doctor's office, exploring issues related to genomic data sharing; the psychosocial effects of direct-to-consumer genetic testing; and the implications of expanded newborn screening for genetic diseases. The field is increasingly focused on the translation of research into broadly accessible applications while promoting individuals' choices about how the information is used and shared for their own care or information.
Lastly, as the promise of the HGP and genomics research to improve health comes closer to being realized, the field continues to engage and excite the public at large. In June, the Smithsonian's National Museum of Natural History in Washington, D.C., in collaboration with NHGRI, will open an exhibition, Genome: Unlocking Life's Code. This exhibition will remain in Washington, D.C., for just over a year and then travel North America for four years. Educational programs associated with the exhibition will include lectures, symposia, discussion panels and gatherings with genomics leaders, scientists, scholars, the arts community and the public.
A decade later, there is still much to learn about the human genome and how to use it to improve the public's health. The availability of personalized genomic information - coupled with rapid changes in how health information is collected, stored, accessed, transferred and used - promises to advance approaches for medical care. Like the HGP, advances en route to an era of genomic medicine require the collective energies and creativities of many researchers, many healthcare professionals and many countries. The genomic revolution is a global effort, with research advances shared around the world.
NHGRI is one of the 27 institutes and centers at the National Institutes of Health. The NHGRI Extramural Research Program supports grants for research and training and career development at sites nationwide. Additional information about NHGRI can be found at http://www.genome.gov.
About the National Institutes of Health (NIH): NIH, the nation's medical research agency, includes 27 institutes and centers and is a component of the U.S. Department of Health and Human Services. NIH is the primary federal agency conducting and supporting basic, clinical, and translational medical research, and is investigating the causes, treatments, and cures for both common and rare diseases. For more information about NIH and its programs, visit http://www.nih.gov.
Contact
Omar McCrimmon
301-402-0911
mccrimmono@mail.nih.gov Posted: April 12, 2013