Last updated: March 14, 2014
Online GWAS Catalog Helps Guide Disease Research
Online GWAS Catalog Helps Guide Disease Research
May 2009
 Researchers who want to sift through the biomedical literature to find genome-wide association results relevant to their research pursuits face an enormous challenge. But, thanks to the efforts of a dedicated team of National Human Genome Research Institute (NHGRI) scientists, they now have an online resource that can make the task a bit less daunting.
Researchers who want to sift through the biomedical literature to find genome-wide association results relevant to their research pursuits face an enormous challenge. But, thanks to the efforts of a dedicated team of National Human Genome Research Institute (NHGRI) scientists, they now have an online resource that can make the task a bit less daunting.
Genome-wide association studies, commonly called GWAS, efficiently scan markers across the DNA, or genomes, of large groups of people looking for variations between individuals with and without a health condition. Over the past few years, this approach has successfully plucked from our DNA code hundreds of the pesky one-letter genetic variations that contribute to the risk of common health conditions, such as obesity and Type 2 diabetes.
The recent deluge of GWAS results coming from these studies is exactly why NHGRI created a resource called A Catalog of Published Genome-Wide Association Studies. The catalog, which has been frequently cited in scientific publications, contains descriptive and association data on hundreds of published common genetic variations and their relationship with nearly 100 complex diseases and traits.
On a weekly basis, epidemiologists from NHGRI's Office of Population Genomics manually curate information from published GWAS and add them to the catalog. Researchers can search the catalog by journal name, first author, disease/trait, statistical significance of association and other categories. Data from the entire catalog can also be directly downloaded as an Excel spreadsheet.
As many had expected, the information in the catalog shows that most genetic variants are relatively common, appearing at a frequency of more than 5 percent in the populations studied. However, each individual genetic variant typically exerts only a very modest effect on risk of each health condition.
The features of the GWAS catalog, as well as an analysis of its contents, are described in a paper published in the May 18 online early edition of the Proceedings of the National Academy of Sciences (PNAS). "Over the last three years, publications describing the results of genome-wide association studies have been pouring out of laboratories," said the paper's senior author, Teri A. Manolio, M.D., Ph.D., director of NHGRI's Office of Population Genomics. "We developed the catalog to track them in a way that might be useful to the research community."
While many genetic variants are often scrutinized individually or in small sets, Dr. Manolio and her colleagues analyzed more than 500 genetic variants from approximately 150 published GWAS to provide clues for the biological function and evolutionary origin of disease-associated regions of the genome.
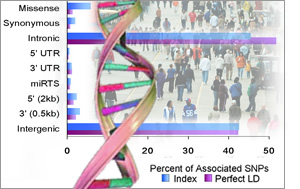
For example, the NHGRI team found that more than 80 percent of the variants that have been associated with a particular trait or disease do not reside within genes, which are the regions of the genome that code for proteins. "Rather, these variants lie in regions of the genome that regulate when, where and to what extent certain genes are turned on," said Praveen Sethupathy, Ph.D., a co-first author of the PNAS study and a postdoctoral fellow in NHGRI's Genome Technology Branch. These findings support recent scientific speculations that non-gene regions of the genome may play a much larger role in disease risk than previously appreciated.
"Our catalog provides a valuable tool for researchers trying to get a handle on what genetic variants have already been found for a particular disease. We hope that this information, along with our recently published analyses, will help to paint a more complete picture of the complex genetic factors involved in common diseases," said Lucia Hindorff, Ph.D., M.P.H., a co-first author of the PNAS study and leader of the catalog team in NHGRI's Office of Population Genomics. "We'll continue to update the catalog and make it widely available to the scientific community as a resource for guiding future research."
Other authors of the PNAS paper include Heather A. Junkins, M.S. and Erin M. Ramos, Ph.D., M.P.H., both of the NHGRI Office of Population Genomics; Jayashri Mehta, M.P.H, an epidemiologist from the National Center for Biotechnology Information, part of the National Library of Medicine; and Francis S. Collins, M.D., Ph.D., former NHGRI director who is a special volunteer conducting research in NHGRI's Genome Technology Branch.