Last updated: July 31, 2012
NIH Expands Human Microbiome Project; Funds Sequencing Centers and Disease Projects

National Human Genome Research Institute
www.genome.gov
NIH Expands Human Microbiome Project; Funds Sequencing Centers and Disease Projects
 Bethesda, Md., Tues., June 23, 2009 — The Human Microbiome Project has awarded more than $42 million to expand its exploration of how the trillions of microscopic organisms that live in or on our bodies affect our health, the National Institutes of Health (NIH) announced today.
Bethesda, Md., Tues., June 23, 2009 — The Human Microbiome Project has awarded more than $42 million to expand its exploration of how the trillions of microscopic organisms that live in or on our bodies affect our health, the National Institutes of Health (NIH) announced today.
The human microbiome is all the microorganisms that reside in or on the human body, as well as all their DNA, or genomes. Launched in 2007 as part of the NIH Common Fund's Roadmap for Medical Research, the Human Microbiome Project is a $140 million, five-year effort that will produce a resource for researchers who are seeking to use information about the microbiome to improve human health.
"This effort will accelerate our understanding of how our bodies and microorganisms interact to influence health and disease," said Acting NIH Director Raynard S. Kington, M.D., Ph.D. "Examining the differences between the microbiomes of healthy patients and those of patients suffering from a disease promises to change how we diagnose, treat and, ultimately, prevent many health conditions."
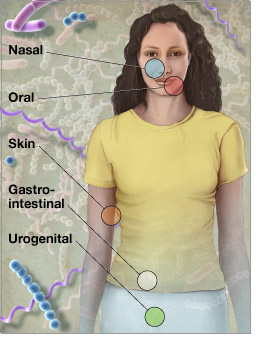
In the new round of funding, the Human Microbiome Project will support the work of the large-scale DNA sequencing centers that participated in the initial phase of the project. These centers will work together to sequence at least 400 microbial genomes. Another approximately 500 microbial genomes are already completed or in sequencing pipelines and supported by individual NIH institutes and internationally funded projects. These data will then be used to characterize the microbial communities found in samples taken from healthy human volunteers. These samples are currently being collected by the Human Microbiome Project from five areas of the body: the digestive tract, the mouth, the skin, the nose and the vagina.
The Human Microbiome Project's large-scale sequencing centers, their principal investigators and approximate funding levels over four years are:
Human Genome Sequencing Center, Baylor College of Medicine, Houston, Richard Gibbs, Ph.D.; $3.7 million
Washington University Genome Sequencing Center, Washington University School of Medicine, Saint Louis, George Weinstock, Ph.D.; $16.1 million
The J. Craig Venter Institute, Rockville, Md., Robert L. Strausberg, Ph.D.; $8.8 million
This center is being funded through the 2009 American Recovery and Reinvestment Act. The ARRA funds that support this center will provide a much needed investment to spur advances in the understanding of how microorganisms that live in or on our bodies affect our health.
In addition, the Broad Institute of MIT/ Harvard, Cambridge, Mass., which participated in the jumpstart phase of the Human Microbiome Project, is expected to participate in this phase of the project.
The Human Microbiome Project is also funding new pilot demonstration projects by researchers that will sample the microbiomes of healthy volunteers and volunteers with specific diseases over the next year. This will allow researchers to study changes in the microbiome at particular body sites that are healthy or affected by diseases. These studies will use samples collected from seven areas of the body: the digestive tract, the mouth, the skin, the nose, the vagina, the blood and the male urethra.
The pilot demonstration projects listed by principal investigator, body site(s), disease focus and approximate funding levels are:
Martin J. Blaser, M.D., New York University School of Medicine
Skin: Psoriasis
$560,000
This study will assess whether changes in the skin microbiome may contribute to psoriasis, an inflammatory skin disease.Gregory A. Buck, Ph.D., Virginia Commonwealth University, Richmond
Vagina: Bacterial Vaginosis and Sexually Transmitted Diseases
$1 million
This study will look at changes in the vaginal microbiome and its association with environmental factors, diseases and a woman's genetic makeup.J. Dennis Fortenberry, M.D., Indiana University Purdue University at Indianapolis
Male urethra: Puberty, Sexual Activity and Sexually Transmitted Diseases
$820,000
Samples will be collected from an ethnically diverse group of male adolescents to characterize the male urethral microbiome and its relation to puberty, sexual activity and sexually transmitted disease.Claire M. Fraser-Liggett, Ph.D., University of Maryland School of Medicine, Baltimore (two grants)
Digestive tract: Obesity
$1 million
This study will sample the gut microbiome from obese and non-obese subjects from the Amish population to uncover associations with metabolism, inflammation, and other traits related to obesity. It will also study the influence of human genetic variation on the composition of the microbiome.Digestive tract: Crohn's Disease
$1 million
The relationship between the inflammatory bowel disease, Crohn's disease, the intestinal microbiome, and bacterial proteins will be explored in samples from pairs of twins with and without the disease.Ellen Li, M.D., Ph.D., Washington University School of Medicine, Saint Louis
Digestive tract: Crohn's Disease
$980,000
This study will analyze the intestinal microbiome and disease subjects' genetic factors to uncover relationships with Crohn's disease, an inflammatory bowel disease.Huiying Li, Ph.D., University of California, Los Angeles
Skin: Acne
$990,000
The goal of this study is to examine the association between the skin microbiome and acne.Zhiheng Pei, M.D., Ph.D., New York University School of Medicine
Mouth and digestive tract: Esophageal Adenocarcinoma
$1 million
This team will sample the oral cavity, esophagus, and stomach to study the relationship of the microbiome from these body sites and esophageal cancer.Jacques Ravel, Ph.D., University of Maryland School of Medicine, Baltimore
Vagina: Bacterial Vaginosis
$980,000
This study will assess the vaginal microbiome and whether it contributes to the risk of bacterial vaginosis, a condition where harmful bacteria outgrow healthy bacteria. Bacterial vaginosis is a risk factor for pre-term delivery and sexually transmitted diseases.Julia Segre, Ph.D., National Human Genome Research Institute, Bethesda, Md.
Skin and nose: Atopic Dermatitis, Immunodeficiency Syndromes
$400,000
The goal of this study is to examine the microbiomes of the skin and nose in three groups of patients with an inflammatory skin disease called atopic dermatitis. Researchers will look for associations between microbiomes, genetic factors associated with atopic dermatitis, and whether patients suffer from immunodeficiency syndromes.Gregory A. Storch, M.D., Washington University School of Medicine, Saint Louis
Nose, digestive tract and blood: Febrile Illness
$950,000
This team of investigators will examine if there is a relationship between the immune system and viral microbiome of groups of children who are healthy and those who are sick with illnesses that cause a sudden high fever.Phillip I. Tarr, M.D., Washington University School of Medicine, Saint Louis
Digestive tract: Necrotizing Enterocolitis
$1 million
This group of researchers will examine the potential connection between the intestinal microbiome and the development of necrotizing enterocolitis, a gastrointestinal disorder that can damage the intestine, in premature infants.James Versalovic, M.D., Ph.D., Baylor College of Medicine, Houston
Digestive tract: Pediatric Irritable Bowel Syndrome (IBS)
$750,000
This project will examine the composition of the intestinal microbiome and any possible connections with irritable bowel syndrome in children.Gary D. Wu, M.D., University of Pennsylvania School of Medicine, Philadelphia
Digestive tract, Crohn's Disease
$1.1 million
Crohn's Disease, a form of inflammatory bowel disease, in children is typically treated with a liquid nutrient diet. It is not known how this treatment works. This team will investigate whether it works by altering the composition of the intestinal microbiome. Partial funding for this project will be provided by NIH's Office of Dietary Supplements.Vincent B. Young, M.D., Ph.D., University of Michigan, Ann Arbor
Digestive tract: Ulcerative Colitis
$1 million
Some patients who suffer from ulcerative colitis, a type of inflammatory bowel disease that causes inflammation and sores in the lining of the rectum and colon, are treated with surgical removal of the colon, followed by construction of a pouch to assist the patient with bowel movements. For unknown reasons, some, patients develop inflammation of the pouch, necessitating further surgery. This research will determine whether the composition of the pouch microbiome influences the onset and progression of pouch inflammation.
Each pilot demonstration project will be reviewed after one year to evaluate its progress toward milestones, as well as its ability to demonstrate a definable relationship between a body site microbiome and disease.
In the first phases of the Human Microbiome Project, jumpstart funding was awarded to create a framework and data resources. Funding has also previously been awarded for the development of innovative technologies and computational tools, coordination of data analysis, and an examination of some of the ethical, legal and social implications of human microbiome research.
More information about the Human Microbiome Project is available at www.nihroadmap.nih.gov/hmp and www.hmpdacc.org.
An illustration showing the body sites that will be sampled as part of the Human Microbiome Project is available at: www.genome.gov/dmd/img.cfm?node=Photos/Graphics&id=85347.
A high resolution image of the bacteria, Entercoccus faecalis, a microbe that lives in the human gut, is available in color at www.genome.gov/dmd/img.cfm?node=Photos/Microorganisms&id=79092, or in black and white at www.genome.gov/dmd/img.cfm?node=Photos/Microorganisms&id=79093.
Some activities described in this release are being funded through the American Recovery and Reinvestment Act (ARRA). More information about NIH's ARRA grant funding opportunities can be found at http://grants.nih.gov/recovery. To track the progress of HHS activities funded through the ARRA, visit www.hhs.gov/recovery. To track all federal funds provided through the ARRA, visit www.recovery.gov.
The Human Microbiome Project is part of the NIH Roadmap for Medical Research funded through the NIH Common Fund and is managed by the National Institute of Allergy and Infectious Diseases, National Institute of Arthritis and Musculoskeletal and Skin Diseases, National Cancer Institute, National Institute of Dental and Craniofacial Research, National Institute of Diabetes and Digestive and Kidney Diseases and National Human Genome Research Institute, and the Office of Strategic Coordination, all part of NIH. The Roadmap is a series of initiatives designed to pursue major opportunities and gaps in biomedical research that no single NIH institute could tackle alone, but which the agency as a whole can address to make the biggest impact possible on the progress of medical research. Additional information about the NIH Roadmap can be found at www.nihroadmap.nih.gov.
The National Institutes of Health (NIH) -"The Nation's Medical Research Agency" - includes 27 Institutes and Centers and is a component of the U. S. Department of Health and Human Services. It is the primary federal agency for conducting and supporting basic, clinical and translational medical research, and it investigates the causes, treatments and cures for both common and rare diseases. For more information about NIH and its programs, visit www.nih.gov.
Contact
Geoff Spencer, NHGRI
301-402-0911
spencerg@mail.nih.gov
NIH Office of Communications
301-496-4461
NIAID News Office
301-402-1663
niaidnews@niaid.nih.gov
Trish Reynolds, NIAMS
301-496-8190
Reynoldsp2@mail.nih.gov
Bob Kuska, NIDCR
301-594-7560
kuskar@nidcr.nih.gov
Leslie Curtis, NIDDK
301-496-3583
niddkmedia@mail.nih.gov