Last updated: September 13, 2013
Genomics and Randomized Trials Network (GARNET)

Background
The Genomics and Randomized Trials Network (GARNET) is series of genome-wide association studies of treatment response in randomized clinical trials. The goal of GARNET is to identify genetic variants associated with response to treatments for conditions of clinical or public health significance. GARNET aims to utilize existing clinical trial data and sample resources to:
- Identify genetic variants that influence an individual's response to treatment.
- Determine whether specific treatments are more or less effective in groups defined by genotype.
- Develop and disseminate innovative methods for adding genome-wide technologies to randomized clinical trials and interpreting the results in the context of a randomized treatment assignment.
Network Structure
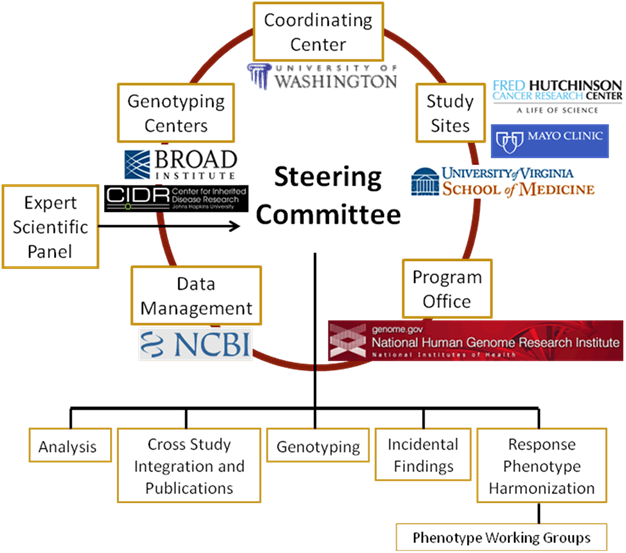
The key extramural participants in GARNET are the study Principal Investigators (PI), the Genotyping Centers and the Coordinating Center. The key National Institutes of Health (NIH) participants are the Office of Population Genomics (OPG) at the National Human Genome Research Institute (NHGRI), which directs the GARNET program, and the National Center for Biotechnology Information (NCBI), which manages the data repository for GARNET studies. GARNET is managed by the participating investigators with substantial cooperation and interaction with NHGRI program staff.
Coordinating Center
The University of Washington (UW) acts as the Coordinating Center and assists with genotypic data cleaning and analysis, phenotype data organization, and coordination of the logistics and administration of the study. The Coordinating Center serves as an internal data repository for GARNET and also assists in phenotypic data harmonization across multiple studies.
Principal Investigator (PI): Bruce Weir, Ph.D.
Co-Principal Investigator (Co-PI): Patrick Heagerty, Ph.D.
Study Investigators
There are three study sites in GARNET:
| Study Investigators | Study Title | Clinical Trial | Phenotypes | Intervention | Study Population Description |
Sample Size |
|---|---|---|---|---|---|---|
| PI: Alex Reiner, M.D., M.Sc. Co-PI: Charles Kooperberg, Ph.D. Fred Hutchinson Cancer Research Center (FHCRC) |
GWAS of Hormone Treatment and CVD and Metabolic Outcomes in the WHI | Women's Health Initiative (WHI) | Vascular events (CHD, stroke, VTE) & Diabetes | Hormone therapy vs. Placebo | 100% female; 100% EA |
5420 |
| PI: Michele Sale, Ph.D. Co-PI: Bradford Worrall, M.D. University of Virginia |
Pharmacogenomic Studies in VISP (Vitamin Intervention for Stroke Prevention) Results & Implications for Clinical Trial Design | Vitamin Intervention for Stroke Prevention (VISP) | Ischemic stroke, MI or death | Ischemic stroke, MI or death/High vs. Low Dose Vitamin therapy (High dose folic acid, vitamins B6 and B12) | 36% female; 82% White, 12% AA | 2,164 |
| PI: Richard Weinshilboum, M.D. Co-PI: Peter Fasching, Ph.D. Mayo Clinic |
A Genome-Wide Association Study in Breast Cancer Patients from the Prospectively Randomized SUCCESS Trial | SUCCESS A | Breast cancer recurrence & cancer associated death | FEC-docetaxel (standard) vs. FEC-docetaxel plus gemcitabine | 100% female; 100% White (1 AA) (Germany) | 3,754 |
Genotyping Centers
There are two laboratories that conduct the genotyping for GARNET studies: the Center for Inherited Disease Research (CIDR) at Johns Hopkins University and the Broad Institute of MIT and Harvard.
Center for Inherited Disease Research (CIDR) at Johns Hopkins University
PI: David Valle, M.D.
Co-PI: Kimberly F. Doheny, Ph.D.
Co-PI: Elizabeth Pugh, Ph.D.
Broad Institute of MIT and Harvard
PI: Stacey Gabriel, Ph.D.
Co-PI: Daniel Mirel, Ph.D.
NIH Participants
National Human Genome Research Institute (NHGRI)
The GARNET program officials at the NHGRI Office of Population Genomics facilitate the achievement of scientific goals and provide institutional oversight and guidance to the project.
National Center for Biotechnology Information (NCBI)
The data from each individual study will be deposited in the Database of Genotypes and Phenotypes (dbGaP), which is housed at NCBI.
Expert Scientific Panel and Steering Committee
The Expert Scientific Panel provides input to the NHGRI director about the progress and direction of the network.
The Steering Committee is responsible for the coordination of all activities conducted by the network.
Steering Committee Chair: Richard Weinshilboum, Mayo Clinic
Subcommittees
GARNET currently has five active subcommittees that meet monthly via teleconference and in-person at Steering Committee meetings in Washington, D.C. Every study has at least one representative on each subcommittee.
The Genotyping Subcommittee, chaired by Kim Doheny, CIDR and Daniel Mirel, Broad Institute, makes recommendations on the thresholds for data cleaning, additional genotyping, and CNV identification and analysis.
The Analysis Subcommittee, chaired by Patrick Heagerty, UW, makes recommendations on cross-study analyses, common issues and analysis elements, develops best practices, and plans future study designs.
The Cross Study Integration & Publications Subcommittee, chaired by Michele Sale, UV, manages and develops policies for cross-study collaboration, publication, presentation, and authorship.
The Incidental Findings Subcommittee, chaired by Brad Worrall, UV, is responsible for developing the GARNET policy for reporting incidental findings and developing recommendations for other studies dealing with these issues.
The Response Phenotype Harmonization Subcommittee, chaired by Alex Reiner, FHCRC and Brad Worrall, UVA, is responsible for identification of common response phenotypes and coordination with other Department of Health and Human Services (DHHS) efforts.
Phenotype Working Groups
For each phenotype category of interest, the Response Phenotype Harmonization Subcommittee is convening a Working Group, led by an investigator from one of the studies, to identify and define specific variables to be shared in cross-study analyses.
GARNET Contacts:
Ebony B. Madden, Ph.D., M.G.C.
Office of Population Genomics, NHGRI
