Contig
Definition
A contig (as related to genomic studies; derived from the word “contiguous”) is a set of DNA segments or sequences that overlap in a way that provides a contiguous representation of a genomic region. For example, a clone contig provides a physical map of a set of cloned segments of DNA across a genomic region, while a sequence contig provides the actual DNA sequence of a genomic region.
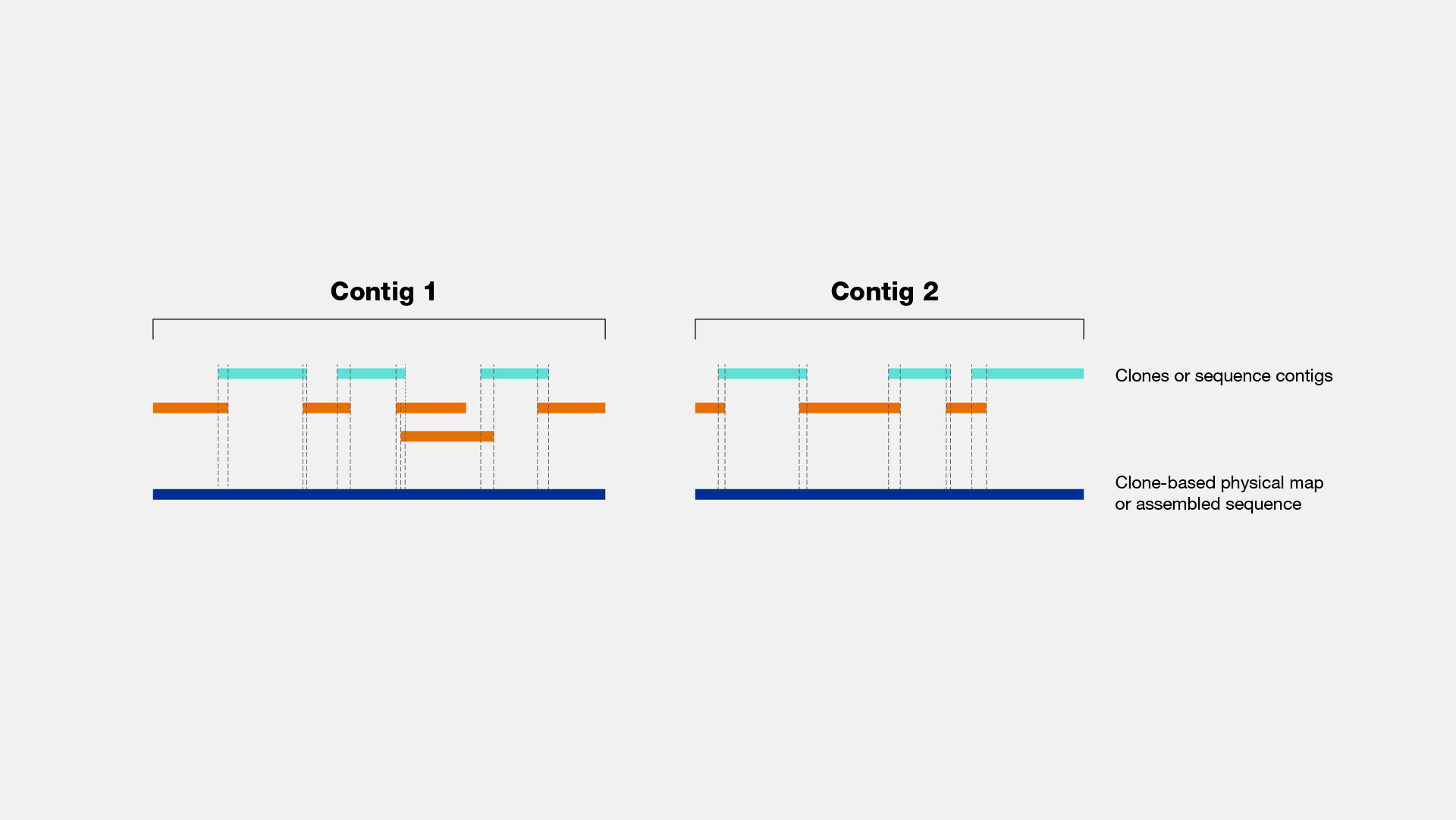
Narration
A genome is typically studied by breaking down its DNA into pieces, analyzing those pieces in the laboratory, and then computationally putting those pieces back together in a conceptual way. Years ago, before genome sequencing became routine, the pieces of DNA would first be cloned and then individual clones would be analyzed to determine which clones overlapped which other clones. A set of overlapping clones that together contain a ‘contiguous’ region of a genome is called a contig – or in this case a clone contig. Genome sequencing uses the concept of a contig, but in a different way. For genome sequencing, the pieces are not actually physical segments of DNA but rather are stretches of generated DNA sequence (often called a sequence read). By establishing the overlaps among a set of sequence reads, one can assemble a sequence contig, from which the actual sequence of the genomic region can be deduced.