Positional Cloning
Definition
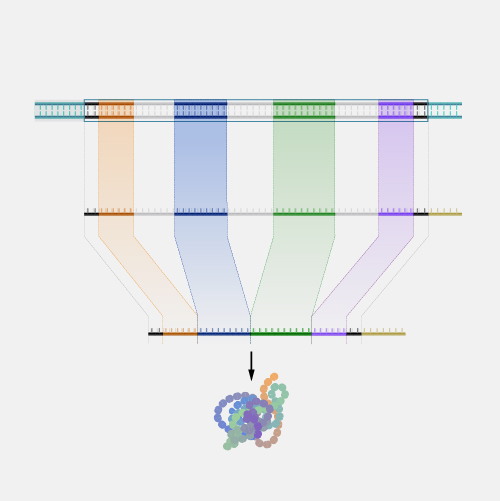
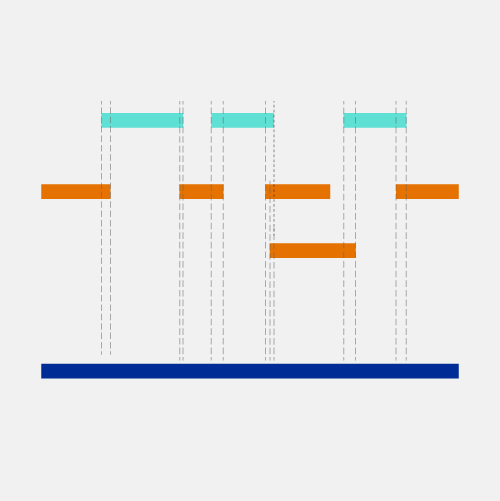
Positional cloning is a laboratory approach used to locate the position of a disease-associated gene on a chromosome. Such a strategy can succeed even when nothing is known about the role of the gene’s encoded protein in the disease. The technique typically relies on the use of known polymorphic markers whose inheritance can be traced through various members of families affected by the disease.
Narration
Positional cloning is a term derived from the late 1980s which basically was to be contrasted to functional cloning, so probably we should define both. Functional cloning was finding a gene by understanding something about what its function is. So the hemophilia gene was identified by knowing there was a problem with a blood clotting factor and then figuring out what gene must have coded for that, and isolating or cloning that gene. But for most diseases, we don't have enough information to guess what the function was, so positional cloning, which came into being as a need of trying to identify the cause of things like cystic fibrosis, was a way of identifying the gene by its position in the genome. Various jokes were made about this. Some people said, oh, positional cloning is a sort of thing where you have to get into a certain position in order to do the work, or if you succeed at it you're guaranteed a position in the university. Not true. Basically, it's the position in the genome that you're trying to zero in on by a series of steps that go from a larger view to narrower view to finally zeroing in on the single base pair that's gone awry. And in many instances that's what you're looking for.