New Genomic Data
The international team of researchers, including those from the National Human Genome Research Institute (NHGRI), made the connection by comparing the whole-genome sequences of 175 ethnic Mongolians to existing genetic variation data.
"Until now, there has been little genomic information from the peoples of North Asia," said Narisu Narisu, Ph.D., bioinformatics scientist at NHGRI. "Our study provides crucial genomic data that fills the information gap for this region and the genomics of present-day East Asians. It potentially will allow Mongolians and related populations to benefit from precision medicine."
The researchers constructed a high quality genetic variation panel that represents people from six Mongolian tribal regions and related populations from Northern Asia. They then incorporated it with data from the 1000 Genomes Project that focuses on Asian populations from Eastern regions.
They discovered that Mongolians share the largest segments of genetic variation inherited from a common ancestor shared with Northern Chinese and Finnish people. Previous studies identified as much as 10.9 percent European ancestry in 10 selected Mongolians currently included in the Human Genome Diversity Panel, while some Northern European populations contained as much as 12 percent Mongolian ancestry.
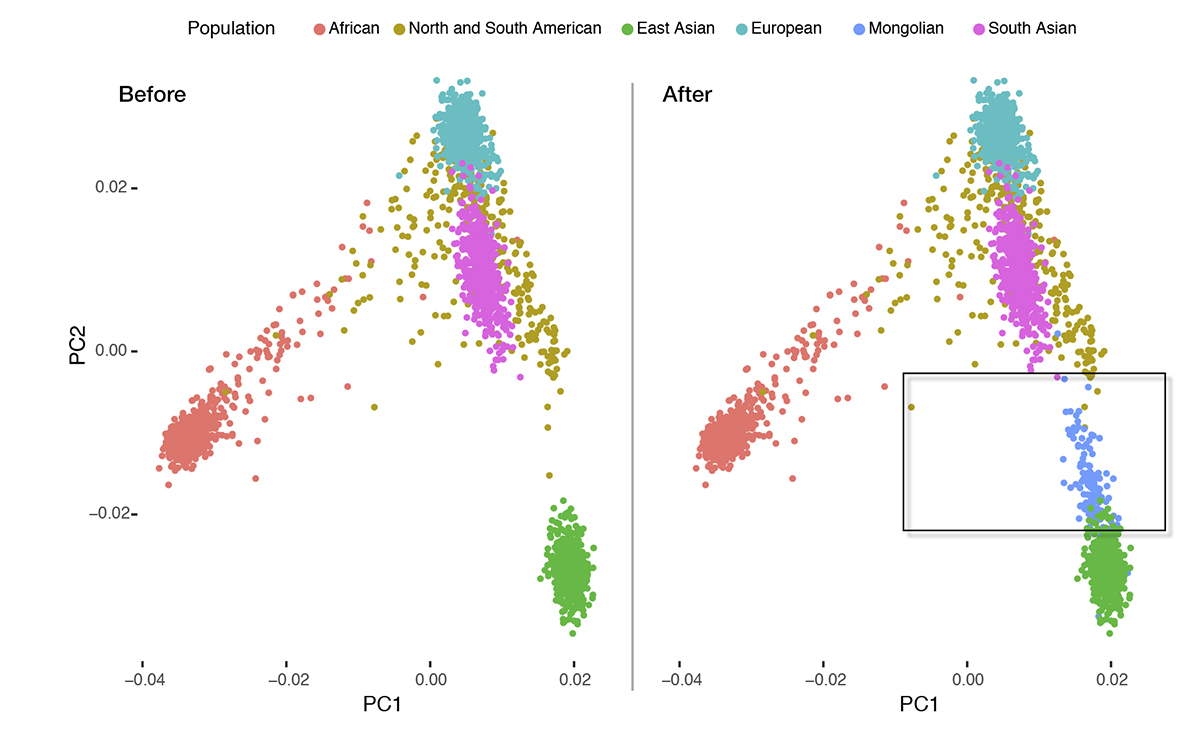
In the new study, researchers also found that East Asians shared more recent common ancestors with Mongolians compared to South Asians. This might indicate that modern East Asians have been isolated from South Asians for long time, and most likely migrated from the ancestors of North Asians.
"Our work may shed light on how humans have adapted to rather harsh geographical conditions and to the nomadic life style of the Mongolian people," said Dr. Narisu, who grew up in the Inner Mongolia Autonomous Region of China. "The data will also help us to study important questions such as origin and migration of East Asian people."
The genetic variation panel could have a potentially huge impact on genomics research: approximately 10 million ethnic Mongolians currently inhabit a wide geographical range that includes Mongolia, Northern China, Southern Russia and other neighboring countries.
Incorporating the study's genetic maps with currently available public data of other ethnic groups will give Dr. Narisu and others a more comprehensive picture of the genomics of human populations. Future studies might include a comparison of the Mongolian genomes with those of Central Asians and less studied, other East Asian groups and a search for the genomic causes of many diseases, in particular those found at higher rates in North Asians.
"We hope the community will use the data from our study to better understand the genomes of all people," said Lawrence Brody, Ph.D., co-author and head of NHGRI's Genetics and Environment Interaction Section. "NHGRI's involvement in this project enhances existing genomic databases. The data are valuable resources for researchers to study diseases and learn how more about the genetic history of Asia."
Historically, NHGRI has supported projects designed to document and understand genomic variation in health and diseases. This includes a number of coordinated large scale sequencing projects such as the HapMap and 1000 Genomes Projects, which specifically sought to sequence the genomes of people from diverse backgrounds.




