NIH-funded startups are fueling the era of genome-completeness
Small businesses with NHGRI grants were instrumental in contributing to the completion of the human genome sequence, redefining the future of genomics.
When Kelvin Liu began his doctorate degree in biomedical engineering at Johns Hopkins University in 2004, he knew his resume was unusual. His previous work painted him as more of an inventor and entrepreneur than an academic. Even before starting his doctoral research, Liu already had two patents under his name. Now, he has eight. Less than two years after graduating with his Ph.D., he became CEO of Circulomics, a biotechnology startup in downtown Baltimore.
According to Liu, the success of Circulomics would not have been possible without support from the National Institutes of Health (NIH) Small Business Innovative Research (SBIR) grants provided by institutes such as the National Human Genome Research Institute (NHGRI), among others.
Small businesses are the heartbeat of innovation in the United States, but they face serious challenges when it comes to raising money to develop their ideas. Front-end money like that provided by NHGRI SBIR grants allows entrepreneurs to focus more on technology building and less on scrambling for early investments.
“It’s not an overstatement to say that SBIR grants from NHGRI and other institutes were both the ignition and gas that propelled Circulomics to where we are today,” Liu said. “Those grants not only acted as a seed but also supported all of our subsequent research and development.”
The NHGRI SBIR grants to Circulomics, totaling approximately $6.8 million, were crucial to catalyze Liu’s science dreams: building technologies that accurately read out the DNA code of all organisms. Federal funding to companies like Liu’s plays a vital role in getting such highly innovative technologies off the ground, fostering the greater U.S. scientific research and development ecosystem.
Until recently, almost all genome sequencing methods required researchers to extract DNA from a given organism and cut the extracted DNA into short pieces. These short pieces were then read out and the resulting sequences strung together in the proper order with the help of specialized computer programs.
This method came with two significant issues. First, the shorter the DNA sequences, the greater the chances that errors could be introduced into the final assembled sequence. Second, when these short DNA sequences are put together, numerous “gaps” emerge — sections where the DNA sequence is too complex to be easily assembled using short snippets.
Many genome scientists have worked tirelessly to develop technologies that would make it easier to start with long DNA pieces and thereby eliminating these two issues.
Since 2014, the NHGRI SBIR grant program has prioritized several areas, including funding researchers who are finding compelling ways to work around the complexities of sequencing complete genomes.
Liu is one of those researchers. Circulomics’ key proprietary technique minimizes the risk of these long DNA strands from breaking. The company developed “Nanobind disks,” which are covered with microscopic silica that binds to DNA and protects it from being damaged.
“It’s a lot like solving a puzzle. If you have 10 big pieces instead of 100 little pieces, assembling gets a lot easier,” said Liu. In 2020, using a DNA sequencing method developed with SBIR funding,Circulomics reported that it had successfully sequenced a 2.44 million base DNA fragment, which at the time was the longest continuous DNA sequence ever generated. Since then, the technology has generated a 4.1 million base DNA sequence, the longest DNA piece ever sequenced (as of publication), according to the company.
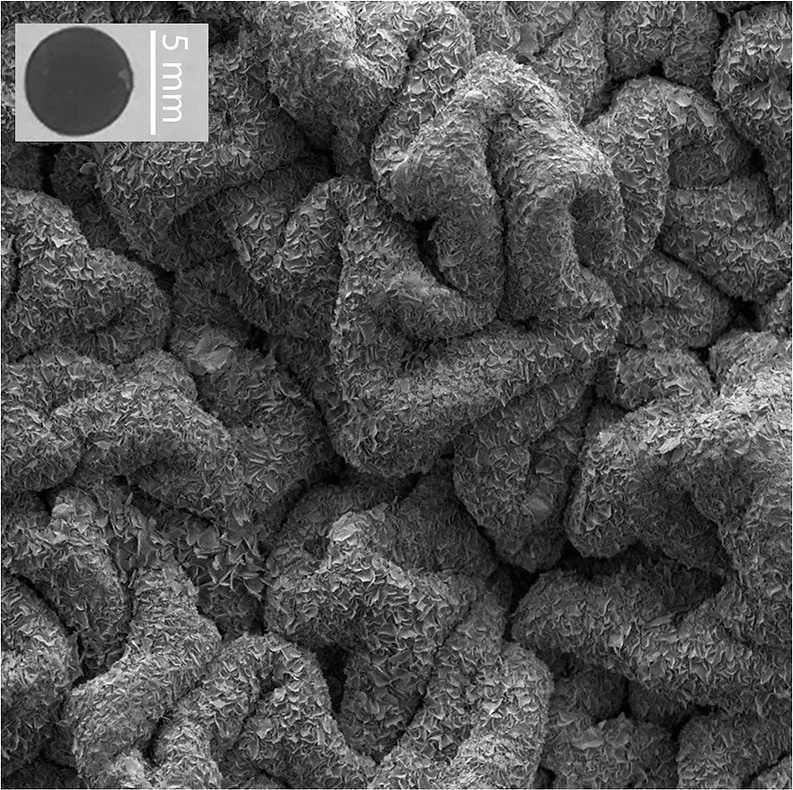

Fig 1. Electron microscope image of Circulomics’ Nanobind disk, formed out of silica. (©Circulomics)
Funding the future
The U.S. Congress passed the Small Business Innovation Development Act of 1982 to spur innovation and help domestic small businesses turn new technologies into viable commercial products. Approximately 3.6% of NHGRI’s budget is now set aside for the program, which amounted to $16 million in 2021. The NIH cumulatively provides funding of over $1 billion each year as part of the SBIR program.
“Small businesses are the heartbeat of innovation in the United States, but they face serious challenges when it comes to raising money to develop their ideas,” said Michael Smith, Ph.D., program director in the NHGRI Genome Technology Program. “Front-end money like that provided by NHGRI SBIR grants allows entrepreneurs to focus more on technology building and less on scrambling for early investments.”
The SBIR funding is unique in that it follows the principle of “non-dilutive equity.” This means that the company does not provide any ownership to the government after it receives the grant. Such no-strings-attached forms of funding allow companies to follow their goals without experiencing any undue burdens.
A bigger picture
When the Human Genome Project ended in 2003, researchers had sequenced approximately 92% of the human genome. The rest of the DNA regions remained unknown because the DNA bases are highly repetitive. Circulomics and other DNA sequencing companies are making it easier to sequence these repeats. But the issue of assembling those pieces back into the larger human genome reference sequence remains.
Repetitive DNA sequences confound researchers abilities to know the number of repeats and their positions in the genome. Even with modern technologies, DNA repeats in the genome are notoriously difficult to study.
DNA is first isolated as multiple fragments and prepared for analysis. Then researchers determine the order of the four bases within each fragment. The result is usually a shattered reconstruction of the genome, with researchers looking at hundreds or thousands of short DNA sequences. Each of these sequences represent a small fragment of a chromosome. Until recently, identifying which sequences were directly next to another (based on the current human genome reference sequence) and getting an accurate picture of more complex regions of the genome were major challenges.
Starting in 2009, a new method called Hi-C changed the genomics landscape. Using Hi-C, researchers finally began to figure out whether two pieces of the genome were next to each other.
“What’s brilliant about Hi-C is that it tells us which pieces go with the same chromosome. If we have a bag of a thousand random pieces, Hi-C tells us that these 10 are from chromosome 1, these 12 are from chromosome 2, and so on,” said Karen Miga, Ph.D., research scientist at the UC Santa Cruz Genomics Institute. “This makes the process of building out a complete genome sequence much easier.”
The NHGRI SBIR program has provided millions of dollars in funding to Arima Genomics and Dovetail Genomics, two major Hi-C companies. Major genome sequencing initiatives frequently use tools and lab kits from both companies.
One such initiative is the Vertebrate Genomes Project, which aims to generate near error-free genome sequences of all vertebrate species. Scientists used Arima’s technologies for the first phase of the project and have successfully sequenced the genomes of 343 species so far.
“Our kits reveal chromosome-level information about the genome as well as interactions between chromosomes,” said Anthony Schmitt, Ph.D., senior vice president at Arima Genomics. “To get such detailed data in a few hours of lab-work — that’s a huge shift.”
Adam Phillippy, Ph.D., is a senior investigator at NHGRI and chair of the Vertebrate Genomes Project’s genome assembly team. He believes using a toolkit that helps determine the genome structure is critical for the project to be successful.
“Every species has a different number of chromosomes, different sizes of chromosomes, and a different way in which the chromosomes are arranged in the nucleus. These technologies provide the basis to study hundreds of species in an affordable and unique way,” said Phillippy.
The search for completeness
After the initial success of the Human Genome Project, the genomics community grappled with the lack of diversity in the human genomes that researchers had sequenced. In 2019, journals, magazines and newspapers across the world began calling for more diverse genomes to be studied.
“This is the power of genomics,” said Sara Hull, Ph.D., director of NHGRI’s Bioethics Core. “By studying the letters of life in all people, and not just some, it inevitably bolsters equity in healthcare and continues the process of building trust with marginalized populations.”
To answer such an imperative, researchers around the globe launched the Human Pangenome Reference Consortium, with extensive NHGRI support. The group is working to generate high-quality, complete human genome sequences representing all ancestries. Circulomics’ longest DNA sequence data are being used as a central part of the consortium.
We tend to think of DNA sequences as being organized in a straight line, which is an incorrect assumption. Each cell in the human body has about three meters of compacted DNA.
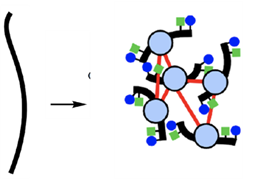
Fig 2. A representative image where pure DNA (left) undergoes Hi-C process, revealing how DNA loops and bundles in a complex manner. Source: Edited for clarity from Green et al., 2015.
“There’s loops, there’s turns, and these chromosomes are interacting with each other in a three-dimensional space.” said Marco Blanchette, Ph.D., vice president of Research and Development at Dovetail Genomics. “We are providing the capacity to accurately build the complete picture of these complex genome organizations.”
The Human Pangenome Reference Consortium recently released sequence data for 30 human genomes. The sequence data were generated using Hi-C kits, including those from Dovetail. By adding new genome sequences from diverse individuals, researchers will be able to identify which regions in the human genome differ among ancestries and whether those regions are associated with human diseases.
Genomicists are hopeful that complete genome sequences and an encyclopedia of genome sequences from people around the globe will provide a foundation for a larger vision. “We are trying to understand human health and disease and building a repository of knowledge for generations to come,” said Smith. “As we start this new chapter, the impact on our knowledge of who we are will be huge.”
Last updated: February 3, 2022
