The Big Picture
- The Human Genome Project was a landmark global scientific effort whose signature goal was to generate the first sequence of the human genome.
- In 2003, the Human Genome Project produced a genome sequence that accounted for over 90% of the human genome. It was as close to complete as the technologies for sequencing DNA allowed at the time.
- The project was critical for advancing policies and earning increased support for the open sharing of scientific data.
- Concerns and questions about sequencing the human genome helped to usher in a greater emphasis on ethics in biomedical research.
- The project was atypical for biomedical research, in that the researchers’ work was driven by a desire to explore an unknown part of the biological world — as opposed to first formulating a theory or hypothesis.

What was the Human Genome Project?
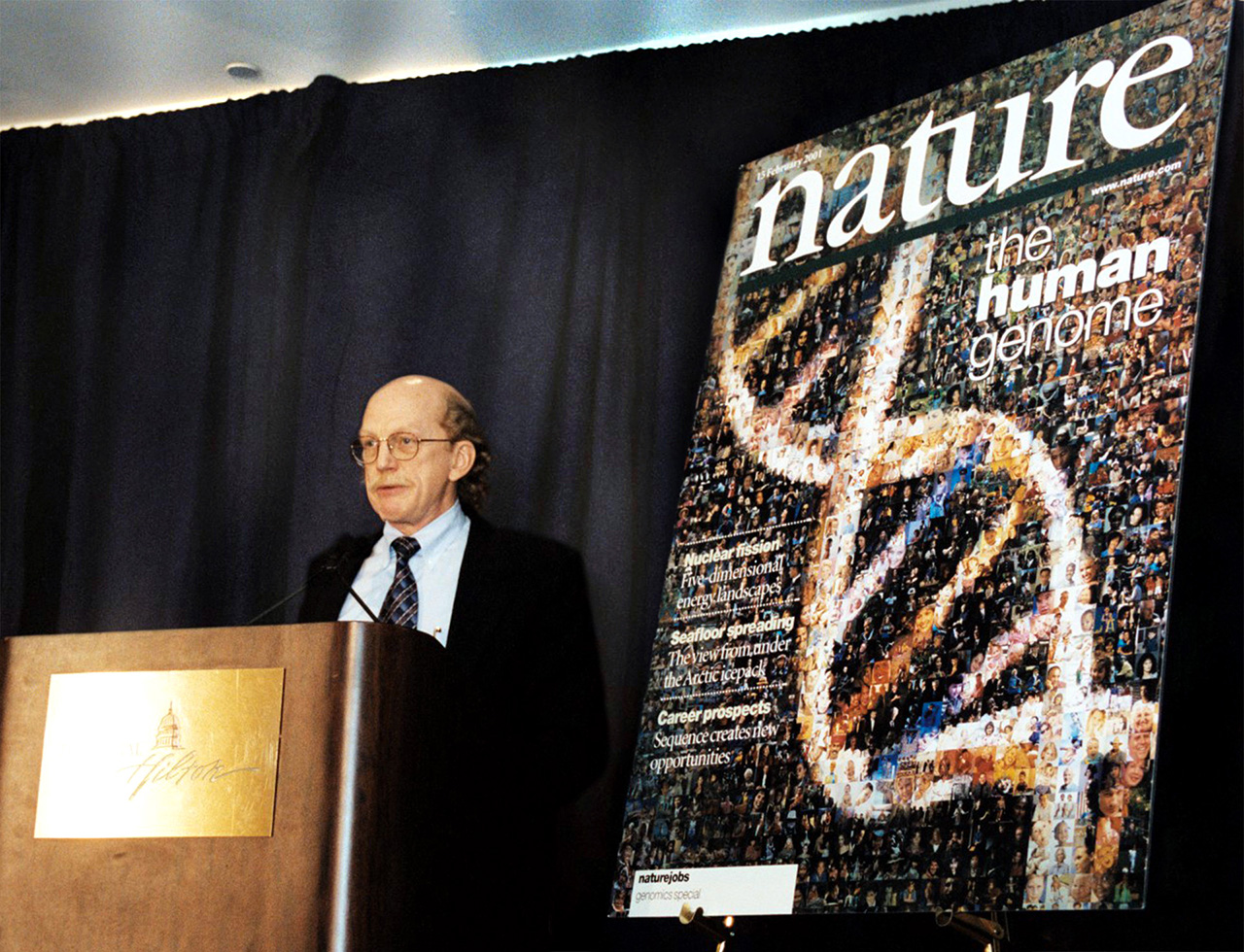
The Human Genome Project was a large, well-organized, and highly collaborative international effort that generated the first sequence of the human genome and that of several additional well-studied organisms. Carried out from 1990–2003, it was one of the most ambitious and important scientific endeavors in human history.
Photo: Robert Waterston, M.D., Ph.D., at the 2001 press conference announcing the publication describing the draft sequence of the human genome generated by the Human Genome Project. Dr. Waterston was an instrumental planner, prominent leader, and major participant of the Human Genome Project. (NHGRI Photo Archive)

What were the goals of the Human Genome Project?
A special committee of the U.S. National Academy of Sciences outlined the original goals for the Human Genome Project in 1988, which included sequencing the entire human genome in addition to the genomes of several carefully selected non-human organisms.
Eventually the list of organisms came to include the bacterium E. coli, baker’s yeast, fruit fly, nematode and mouse. The project’s architects and participants hoped the resulting information would usher in a new era for biomedical research, and its goals and related strategic plans were updated periodically throughout the project.
In part due to a deliberate focus on technology development, the Human Genome Project ultimately exceeded its initial set of goals, doing so by 2003, two years ahead of its originally projected 2005 completion. Many of the project’s achievements were beyond what scientists thought possible in 1988.
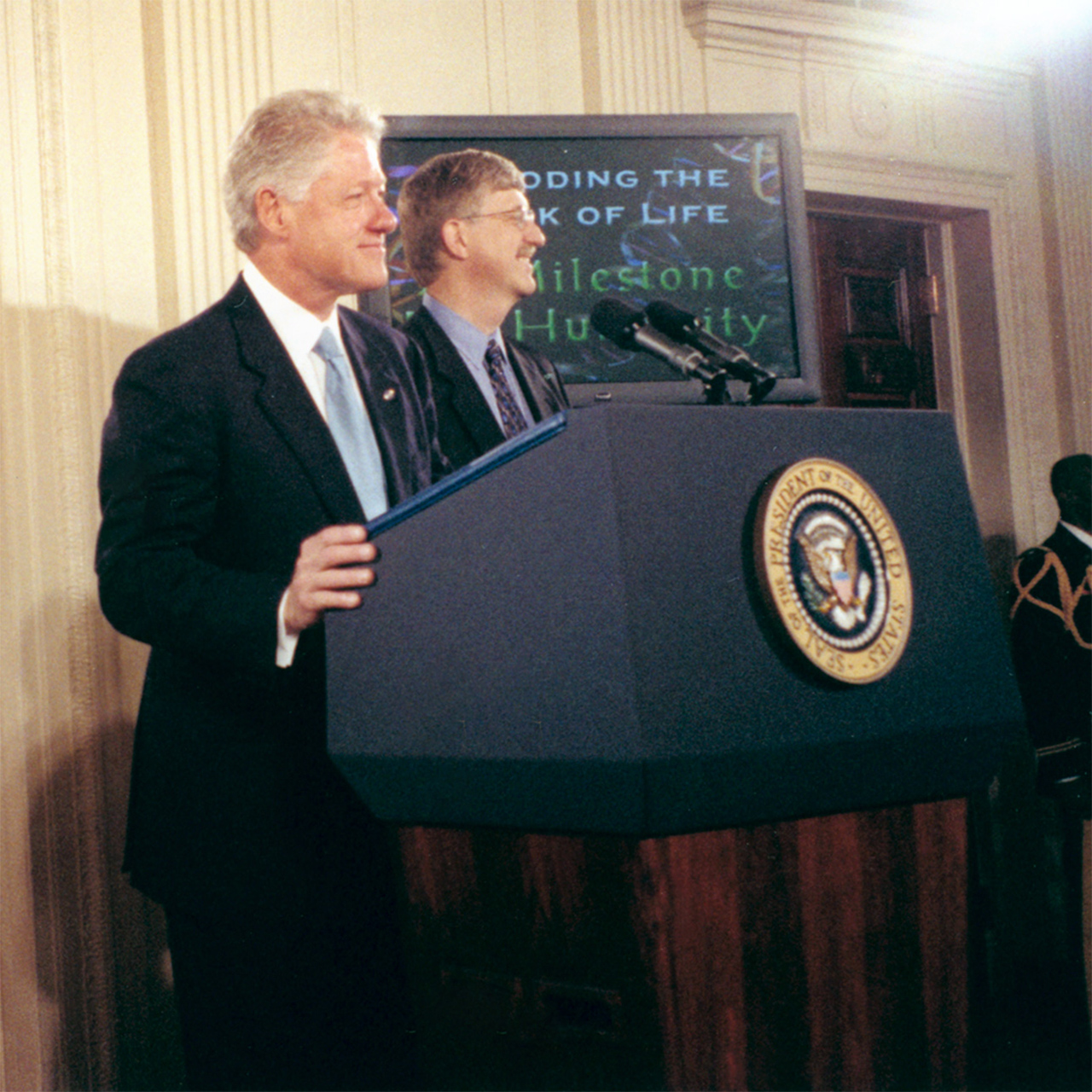
President Bill Clinton and Francis Collins, M.D., Ph.D., (NHGRI Director) at a June 2000 event at the White House celebrating the draft human genome sequence generated by the Human Genome Project. Dr. Collins served as the de facto leader of the International Human Genome Sequencing Consortium, the group that sequenced the human genome during the Human Genome Project. (NHGRI Photo Archive)

What is DNA sequencing? How was it performed during the Human Genome Project?
DNA sequencing involves determining the exact order of the bases in DNA — the As, Cs, Gs and Ts that make up segments of DNA. Because the Human Genome Project aimed to sequence all of the DNA (i.e., the genome) of a set of organisms, significant effort was made to improve the methods for DNA sequencing.
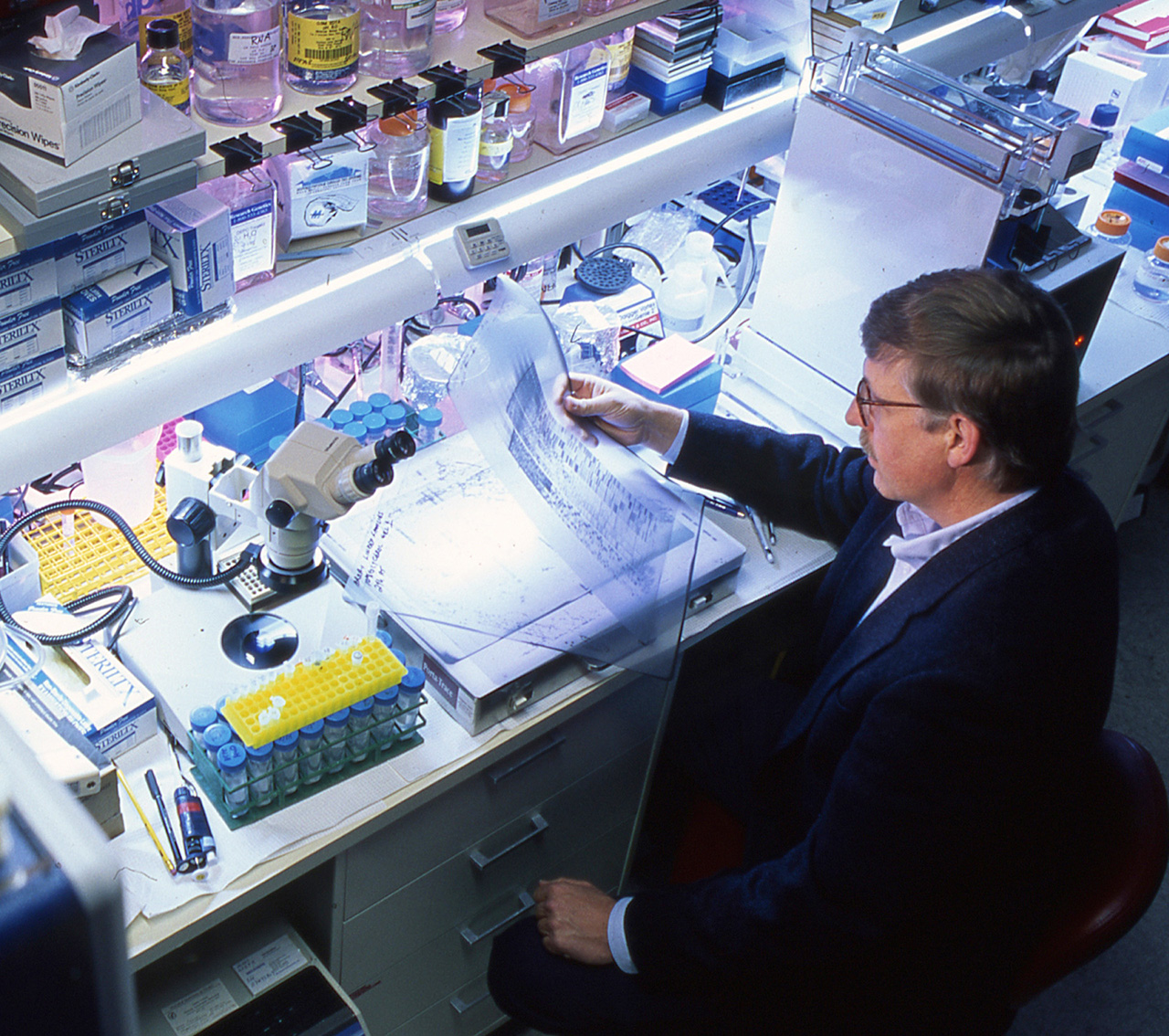
Ultimately, the project used one particular method for DNA sequencing, called Sanger DNA sequencing, but first greatly advanced this basic method through a series of major technical innovations.
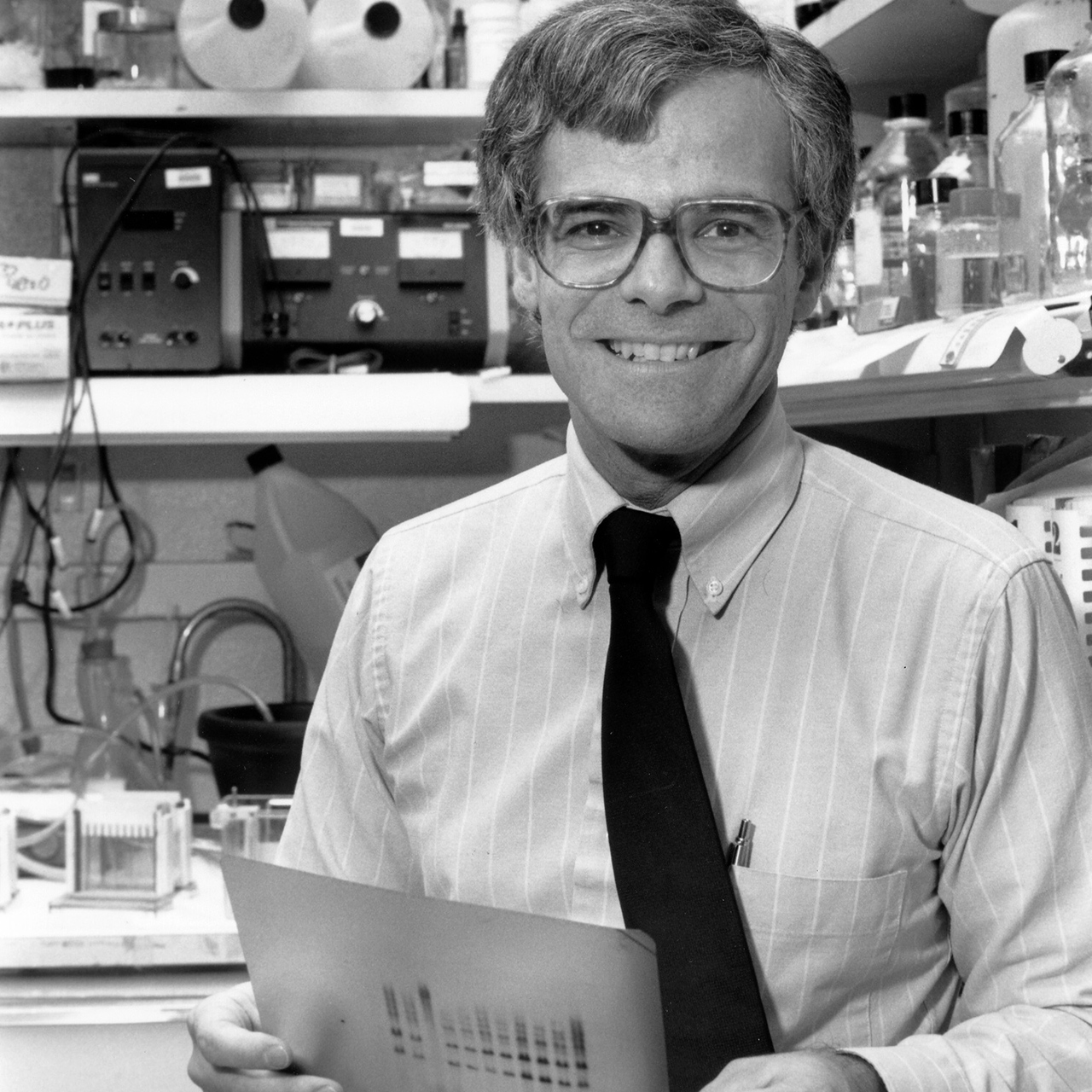
Photo: Dr. Collins analyzing an autoradiogram displaying the results of a Sanger DNA sequencing experiment, such as that used in the early years of the Human Genome Project. (NHGRI Photo Archive)
Whose genome was sequenced by the Human Genome Project?
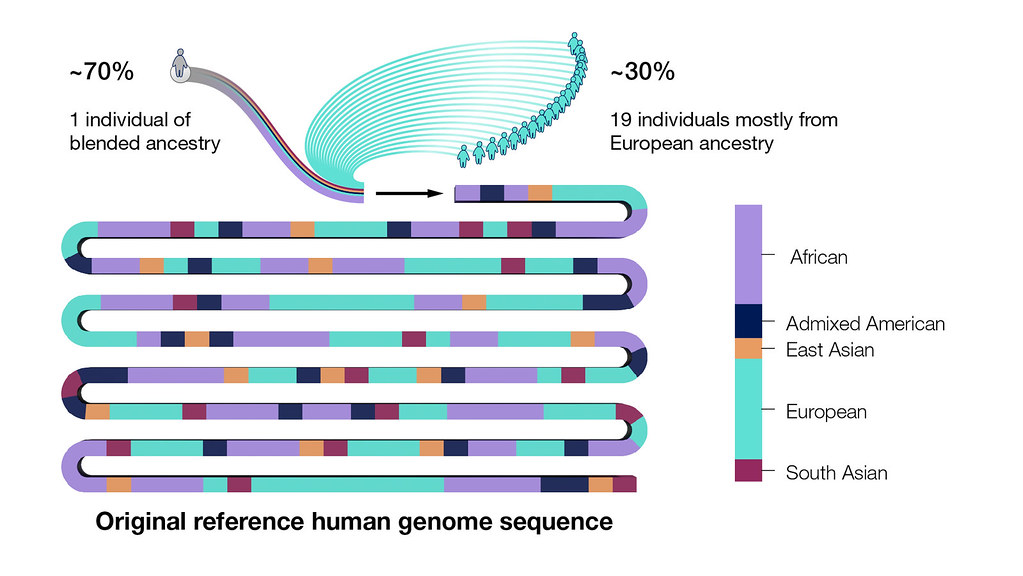
The human genome sequence generated by the Human Genome Project was actually a patchwork of multiple people whose identities were intentionally made anonymous to protect their privacy.
However, the majority of the sequence came from one person of blended ancestry. Specifically, 70 percent of the original reference human genome sequence was generated from that individual’s DNA, with the remaining 30 percent coming from a combination of 19 other individuals of mostly European ancestry.

How was the DNA collected by the Human Genome Project?
Most of the original human genome sequence came from volunteers living in Buffalo, New York. Researchers at the Roswell Park Cancer Institute, located in Buffalo, were experts at preparing the DNA in a form that could be used for sequencing the human genome
The researchers recruited volunteer donors through public advertisements (like the one on the right). They then acquired informed consent from these volunteers and collected blood samples, from which they extracted DNA.
Photo: A Buffalo, New York, newspaper advertisement from 1997 recruiting volunteers to provide blood samples and DNA for the Human Genome Project. (NHGRI History of Genomics Program Archive)

Who carried out the Human Genome Project?
The Human Genome Project could not have been completed as quickly and effectively without the dedicated participation of an international consortium of thousands of researchers. In the United States, the researchers were funded by the Department of Energy and the National Institutes of Health, which created the Office for Human Genome Research in 1988 (later renamed the National Center for Human Genome Research in 1990 and then the National Human Genome Research Institute in 1997).
The sequencing of the human genome involved researchers from 20 separate universities and research centers across the United States, United Kingdom, France, Germany, Japan and China. The groups in these countries became known as the International Human Genome Sequencing Consortium.
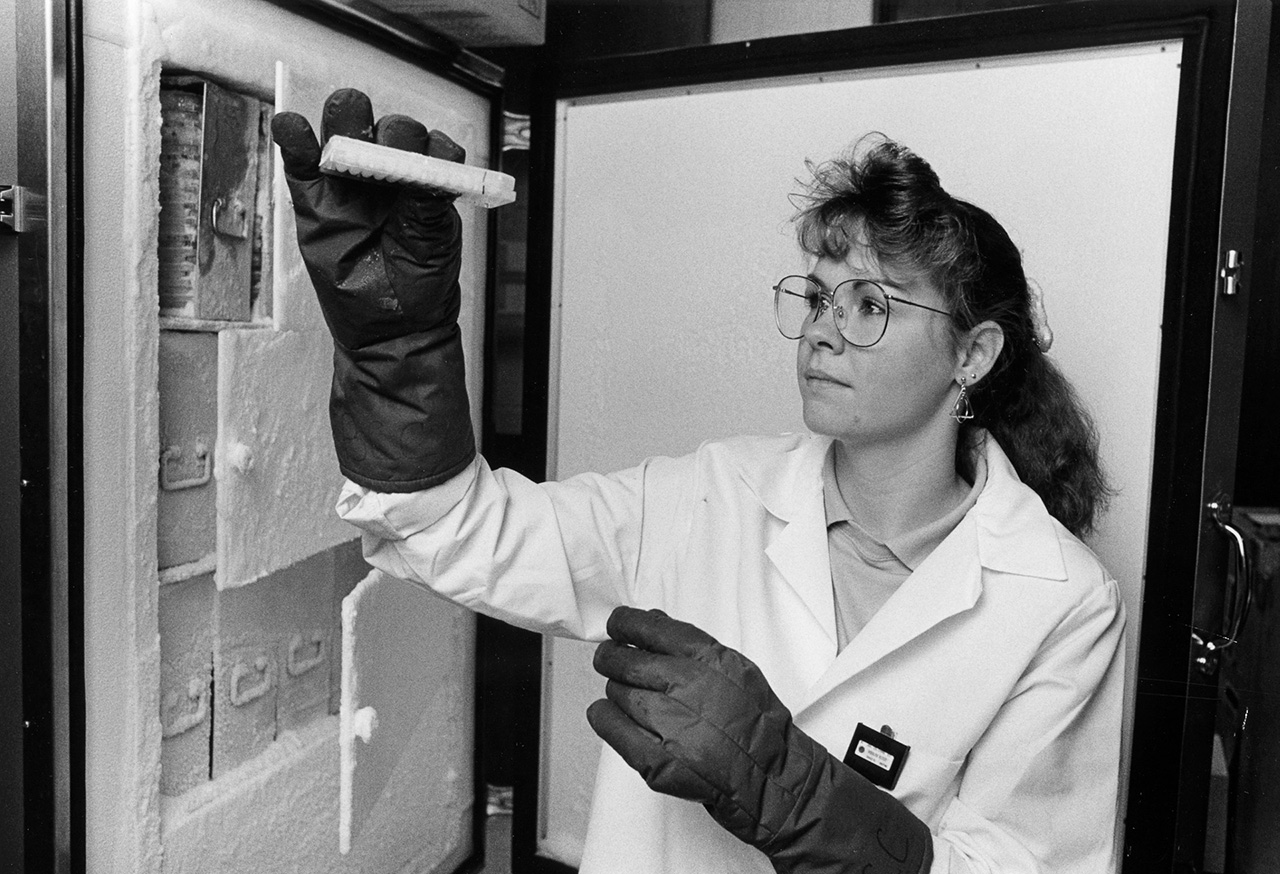
Photo: Researcher at Washington University in St. Louis handling frozen clones containing human DNA being studied by Human Genome Project researchers. (NHGRI Photo Archive)

How much did the Human Genome Project cost?
The initially projected cost for the Human Genome Project was $3 billion, based on its envisioned length of 15 years. While precise cost-accounting was difficult to carry out, especially across the set of international funders, most agree that this rough amount is close to the accurate number.
The cost of the Human Genome Project, while in the billions of dollars, has been greatly offset by the positive economic benefits that genomics has yielded in the ensuing decades. Such economic gains reflect direct links between resulting products and advances in the pharmaceutical and biotechnology industries, among others.
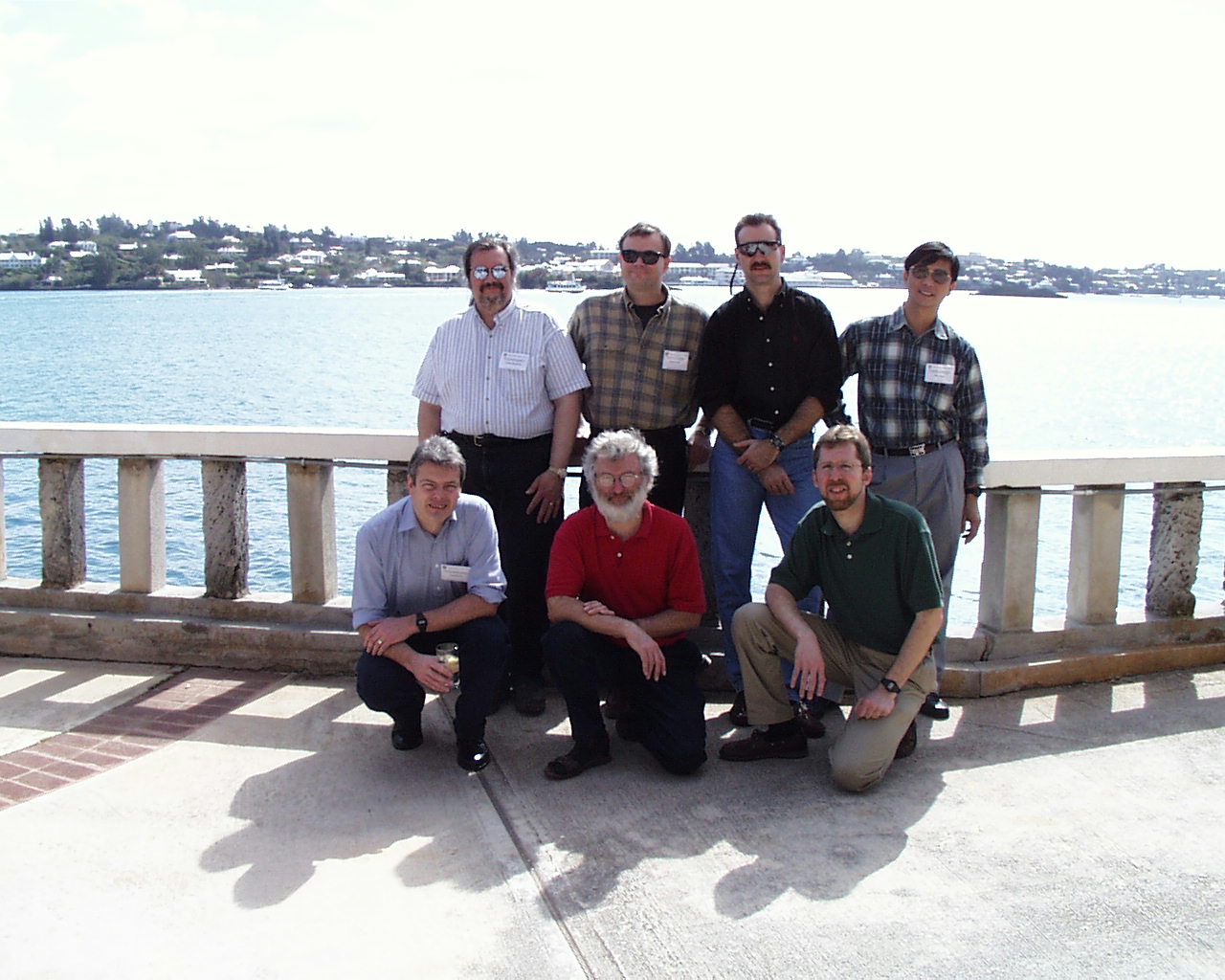
Photo: A subset of the participants of a 1996 meeting held in Bermuda that helped to establish the “Bermuda Principles” for sharing genome sequence data during the Human Genome Project. Shown (from left to right) are: David Bentley, John Sulston, and Eric Green in the front row and Richard McCombie, Richard Gibbs, Richard Wilson, and Elson Chen in the back row. (Photo by Eric Green)

Did the Human Genome Project produce a perfectly complete genome sequence?
No. Throughout the Human Genome Project, researchers continually improved the methods for DNA sequencing. However, they were limited in their abilities to determine the sequence of some stretches of human DNA (e.g., particularly complex or highly repetitive DNA).
In June 2000, the International Human Genome Sequencing Consortium announced that it had produced a draft human genome sequence that accounted for 90% of the human genome. The draft sequence contained more than 150,000 areas where the DNA sequence was unknown because it could not be determined accurately (known as gaps).
In April 2003, the consortium announced that it had generated an essentially complete human genome sequence, which was significantly improved from the draft sequence. Specifically, it accounted for 92% of the human genome and less than 400 gaps; it was also more accurate.
On March 31, 2022, the Telomere-to-Telomere (T2T) consortium announced that had filled in the remaining gaps and produced the first truly complete human genome sequence.
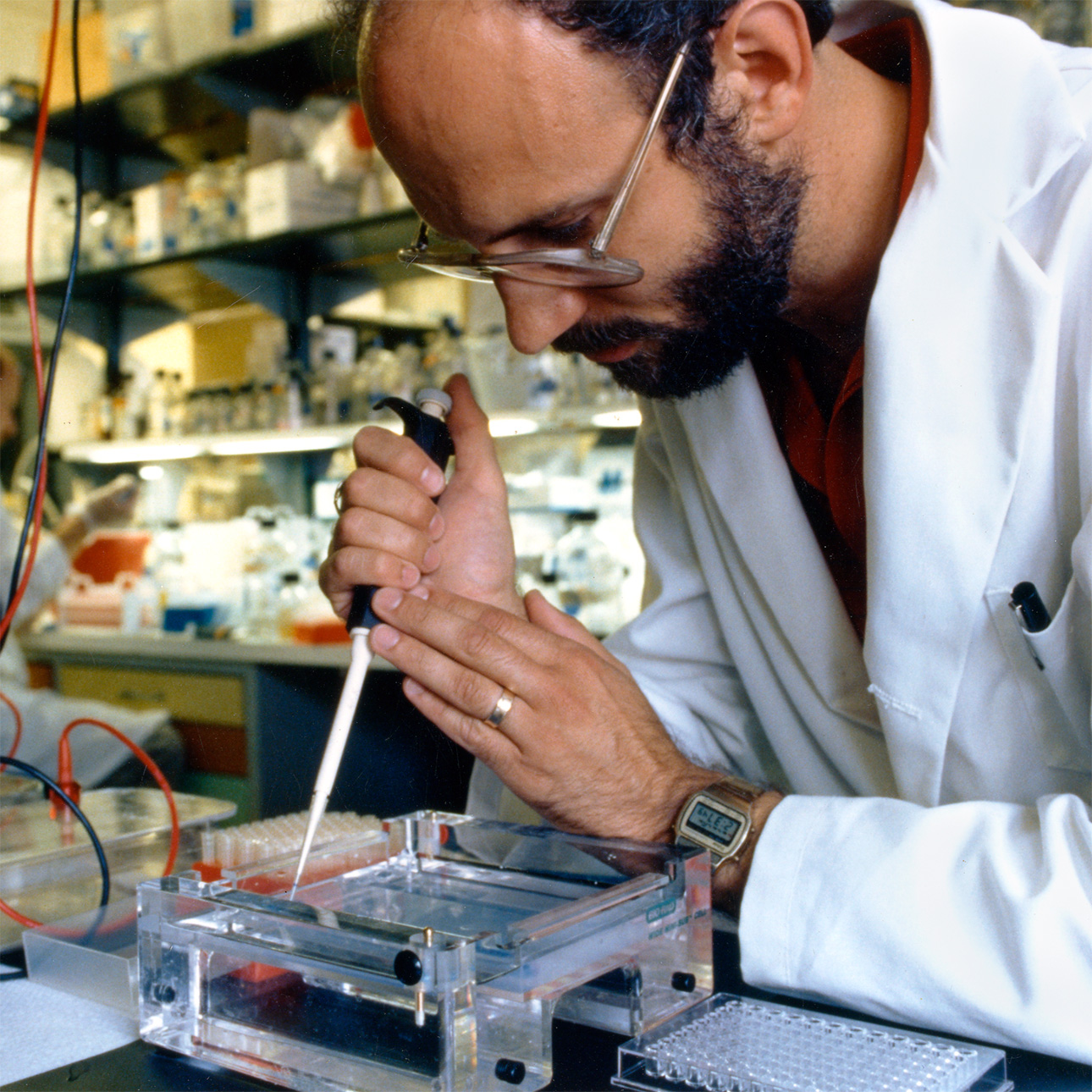
Photo: A Human Genome Project researcher pipetting a DNA sample into an agarose gel to perform gel electrophoresis. (NHGRI Photo Archive)

How did the Human Genome Project change practices around data sharing in the scientific research community?
Human Genome Project scientists made every part of the draft human genome sequence publicly available shortly after production.
This routine came from two meetings in Bermuda in which project researchers agreed to the “Bermuda Principles,” which set out the rules for the rapid release of sequence data. This landmark agreement has been credited with establishing a greater awareness and openness to the sharing of data in biomedical research, making it one of the most important legacies of the Human Genome Project.
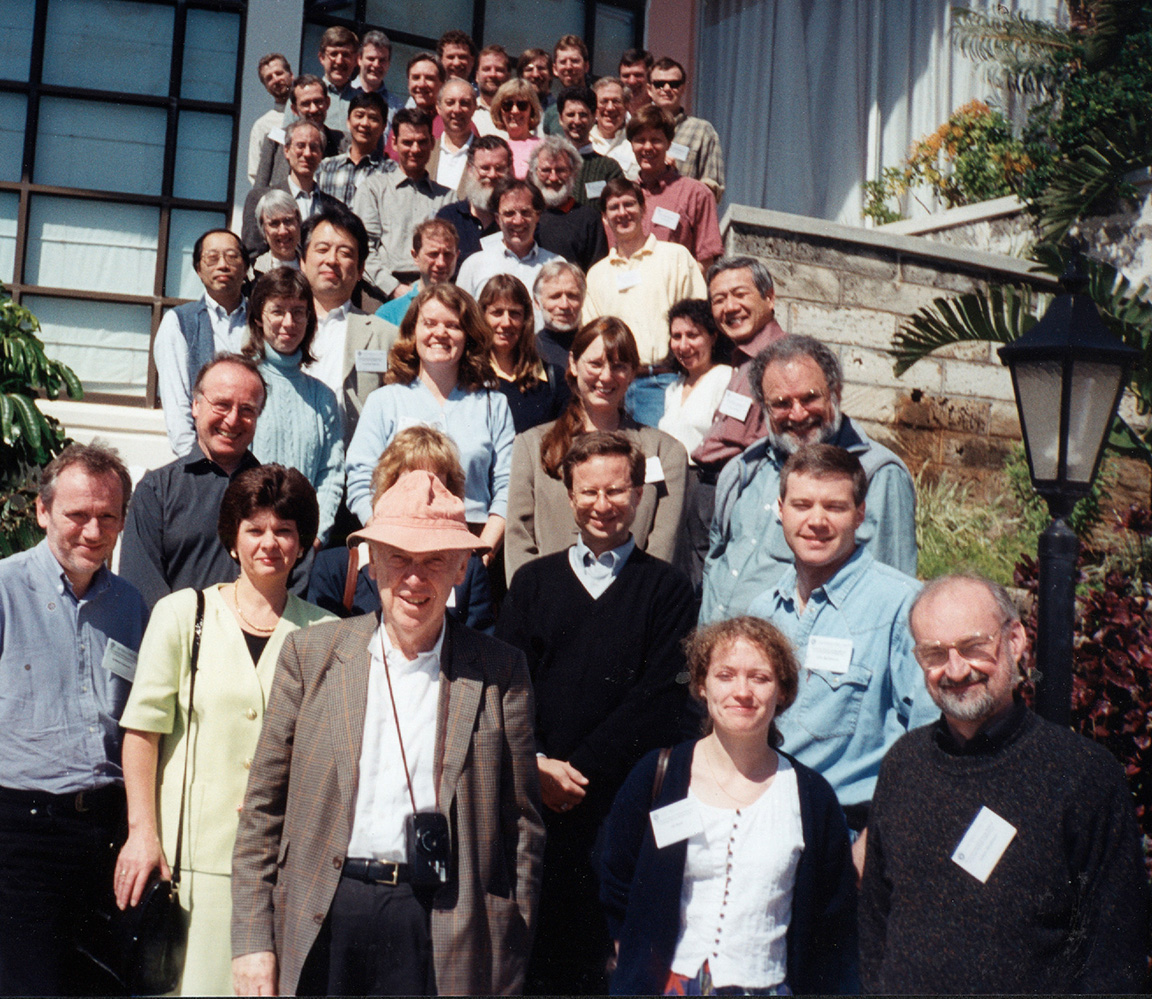
The first International Strategy Meeting on Human Genome Sequencing in Bermuda 1996. Nobelist James Watson, Ph.D., (front left; in hat) played an instrumental role in launching the Human Genome Project and promoting the project’s sharing of genomic data; however, Dr. Watson is also known for his offensive and scientifically incorrect comments about a number of societal topics that are incongruent with the values of NHGRI. (NHGRI Photo Archive)

How did the Human Genome Project foster ethics in biological research?
The leaders of the Human Genome Project recognized the need to be proactive in addressing a wide range of ethical and social issues related to the acquisition and use of genomic information. They were especially aware of the potential risks and benefits of incorporating new genomic knowledge into research and medicine. Similarly, they were aware of the potential misuse of genomic information when it came to insurance and employment, among others.
To help understand and address these issues, NHGRI established the Ethical, Legal, and Social Implications (ELSI) Research Program in 1990.
The early appreciation of the value of this program later led the U.S. Congress to mandate that NHGRI dedicate at least 5% of its research budget to studying the ethical, legal and social implications of genomic advances. The NHGRI ELSI Research Program has become a model for bioethics research worldwide.
Former NHGRI Acting Director Michael Gottesman, M.D., during the Human Genome Project. Dr. Gottesman played an important interim role in leading NHGRI between its first Director (Dr. Watson) and its second Director (Dr. Collins). (NHGRI Photo Archive)
How did the Human Genome Project affect biological research in general?
The Human Genome Project demonstrated that production-oriented, discovery-driven scientific inquiry—which did not involve the investigation of a specific hypothesis or the direct answering of preformed questions—could be remarkably valuable and beneficial to the broader scientific community.
The project was also a successful example of “big science” in biomedical research. The magnitude of the technological challenges prompted the Human Genome Project to assemble interdisciplinary groups from across the world, involving experts in engineering, biology, and computer science, among other areas. It also required the work to be concentrated in a modest number of major centers to maximize economies of scale.
Before the Human Genome Project, the biomedical research community viewed projects of such scale with deep skepticism. These kinds of massive scientific undertakings have become more commonplace and well-accepted based in part on the success of the Human Genome Project.
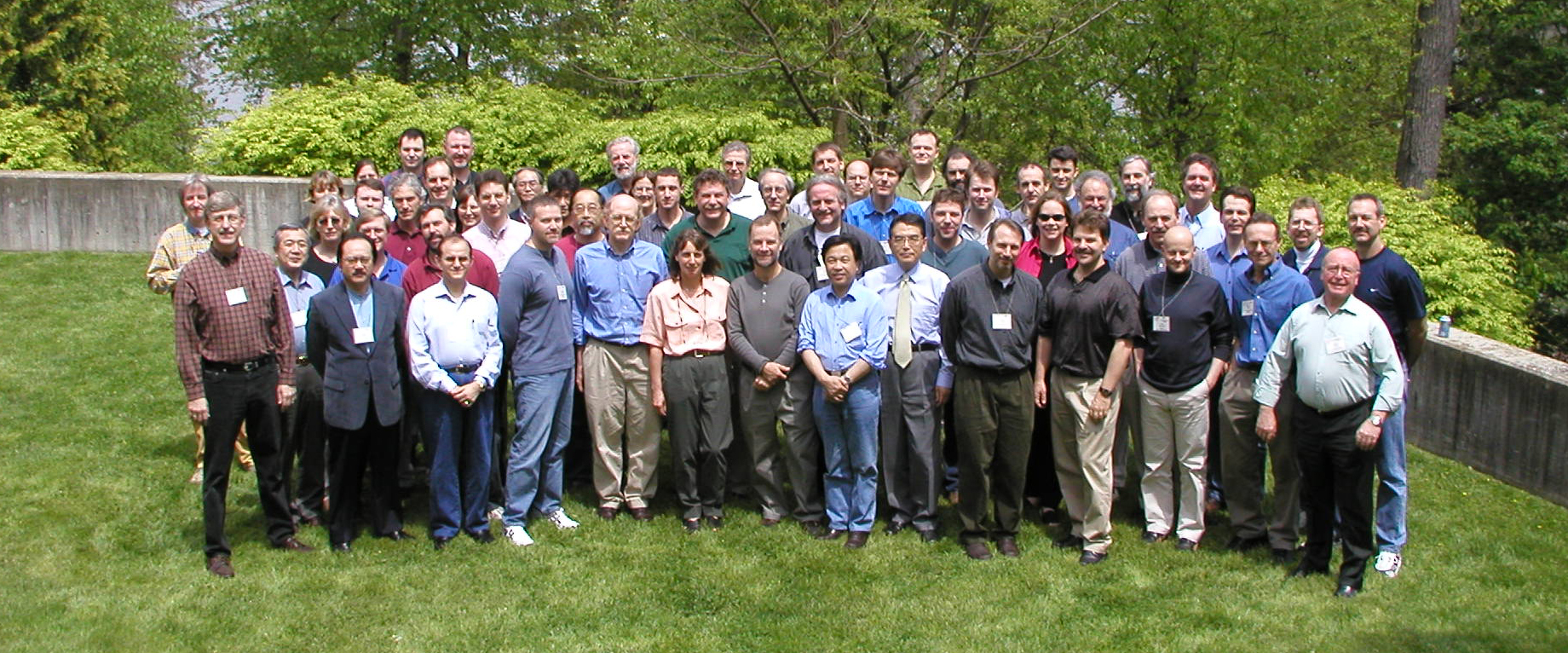

Caption: Participants of a 2002 meeting (held at Cold Spring Harbor Laboratory) of the International Human Genome Sequencing Consortium, the group involved in generating the first human genome sequence during the Human Genome Project. Credit: Eric Green, NHGRI.
Last updated: June 13, 2024
