Comparative Genomics Fact Sheet
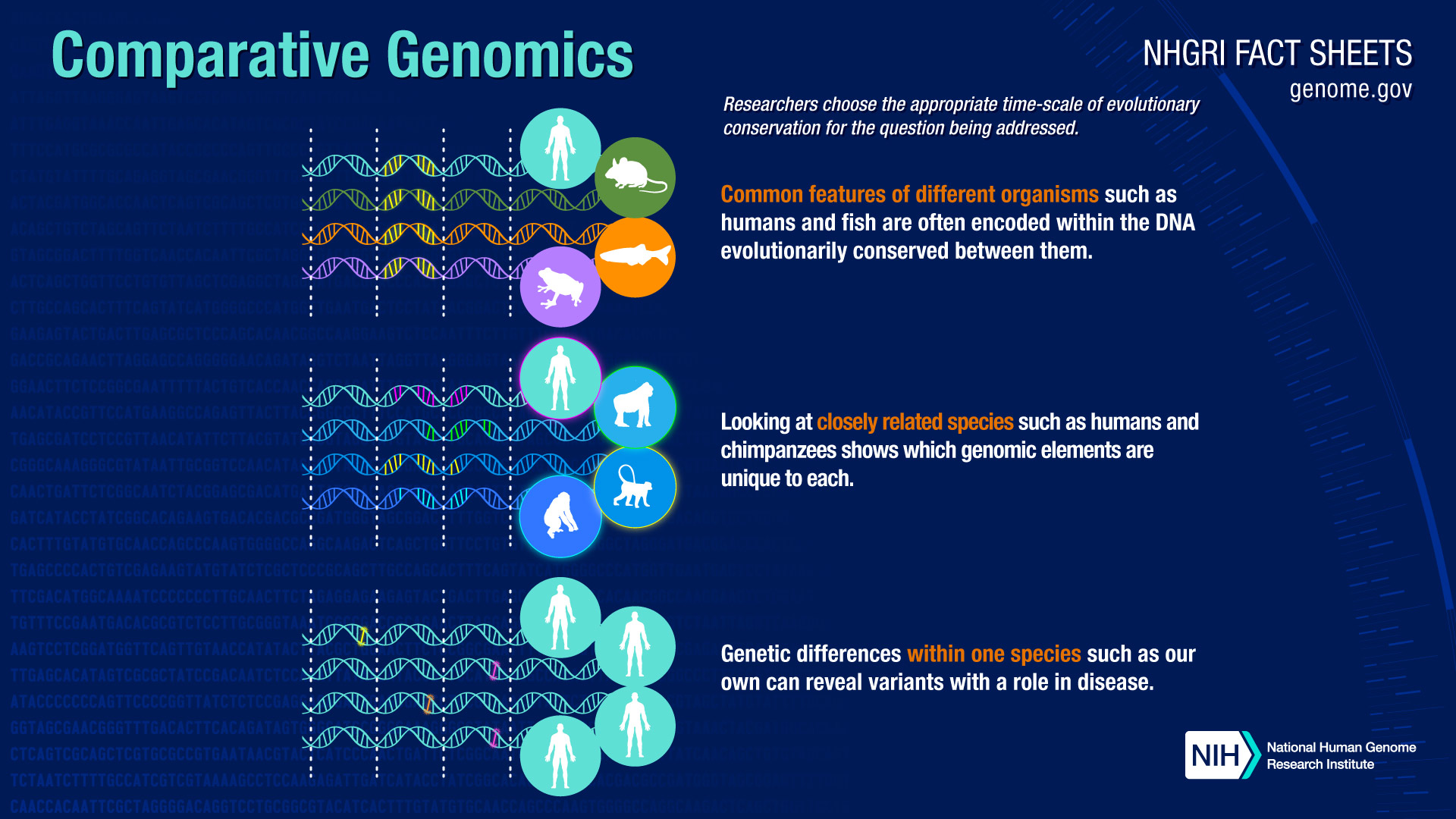
Comparative genomics is a field of biological research in which researchers use a variety of tools to compare the complete genome sequences of different species. By carefully comparing characteristics that define various organisms, researchers can pinpoint regions of similarity and difference.
What are the benefits of comparative genomics?
Identifying DNA sequences that have been "conserved" - that is, preserved in many different organisms over millions of years - is an important step toward understanding the genome itself. It pinpoints genes that are essential to life and highlights genomic signals that control gene function across many species. It helps us to further understand what genes relate to various biological systems, which in turn may translate into innovative approaches for treating human disease and improving human health.
Comparative genomics also provides a powerful tool for studying evolution. By taking advantage of - and analyzing- the evolutionary relationships between species and the corresponding differences in their DNA, scientists can better understand how the appearance, behavior and biology of living things have changed over time.
As DNA sequencing technology becomes more powerful and less expensive, comparative genomics is finding wider applications in agriculture, biotechnology and zoology as a tool to tease apart the often subtle differences among animal species. Such efforts have led to new insights into some branches on the evolutionary tree, as well as improving the health of domesticated animals and pointing to new strategies for conserving rare and endangered species.
-
What are the benefits of comparative genomics?
Identifying DNA sequences that have been "conserved" - that is, preserved in many different organisms over millions of years - is an important step toward understanding the genome itself. It pinpoints genes that are essential to life and highlights genomic signals that control gene function across many species. It helps us to further understand what genes relate to various biological systems, which in turn may translate into innovative approaches for treating human disease and improving human health.
Comparative genomics also provides a powerful tool for studying evolution. By taking advantage of - and analyzing- the evolutionary relationships between species and the corresponding differences in their DNA, scientists can better understand how the appearance, behavior and biology of living things have changed over time.
As DNA sequencing technology becomes more powerful and less expensive, comparative genomics is finding wider applications in agriculture, biotechnology and zoology as a tool to tease apart the often subtle differences among animal species. Such efforts have led to new insights into some branches on the evolutionary tree, as well as improving the health of domesticated animals and pointing to new strategies for conserving rare and endangered species.
What is a genome made of?
The genomes of almost all living creatures, both plants and animals, consist of DNA (deoxyribonucleic acid), the chemical chain that includes the genes that code for different proteins and the regulatory sequences that turn those genes on and off. Precisely which protein is produced by any given gene is determined by the sequence in which four building blocks - adenine (A), thymine (T), cytosine (C) and guanine (G) - are laid out along DNA's twisted, double-helix structure.
-
What is a genome made of?
The genomes of almost all living creatures, both plants and animals, consist of DNA (deoxyribonucleic acid), the chemical chain that includes the genes that code for different proteins and the regulatory sequences that turn those genes on and off. Precisely which protein is produced by any given gene is determined by the sequence in which four building blocks - adenine (A), thymine (T), cytosine (C) and guanine (G) - are laid out along DNA's twisted, double-helix structure.

What results has the field of comparative genomics produced?
Comparative genomics has yielded dramatic results. Investigators are increasingly using comparative genomics to explore areas ranging from human development and behavior to metabolism and susceptibility to disease. These studies are uncovering new behavioral, neurological and developmental pathways and genes that are shared or related among species. Some researchers are using comparative genomics to reveal the genomic underpinnings of disease in animals with the hope of gaining new insights into disease development in humans.
Among the results so far are the following:
-
A study discovered that about 60 percent of genes are conserved between fruit flies and humans, meaning that the two organisms appear to share a core set of genes. Two-thirds of human genes known to be involved in cancer have counterparts in the fruit fly.
-
A comparative genomics analysis of six species of yeast prompted scientists to significantly revise their initial catalog of yeast genes and to predict a new set of functional elements that play a role in regulating genome activity, not just in yeast but across many species.
-
Researchers studying milk production have mapped genes that increase the yield of high-fat milk in cows, resulting in higher production levels and potentially a significant economic impact. This is one of many studies aimed at increasing food production.
-
Scientists have found genes that increase muscling in cattle by twofold; they found the same genes in racing dogs, and such results may foster human performance studies.
-
Comparisons of nearly 50 bird species' genomes revealed a gene network that underlies singing in birds and that may have an important role in human speech and language. The bird researchers also found gene networks responsible for traits such as feathers and beaks.
-
In recent years, researchers in the National Human Genome Research Institute (NHGRI) intramural program also have studied the genomics of various cancer types in dogs, including common cancers and other diseases, to try to develop new insights into the human form of the condition. In some cases, they have mapped genes contributing to these disorders.
-
In other studies, NHGRI researchers are comparing how genes affect body shape and size in dogs to better understand growth and development. Studies of dogs with sleep problems have revealed genes and pathways - and potential drug targets - to treat sleep problems.
-
What results has the field of comparative genomics produced?
Comparative genomics has yielded dramatic results. Investigators are increasingly using comparative genomics to explore areas ranging from human development and behavior to metabolism and susceptibility to disease. These studies are uncovering new behavioral, neurological and developmental pathways and genes that are shared or related among species. Some researchers are using comparative genomics to reveal the genomic underpinnings of disease in animals with the hope of gaining new insights into disease development in humans.
Among the results so far are the following:
-
A study discovered that about 60 percent of genes are conserved between fruit flies and humans, meaning that the two organisms appear to share a core set of genes. Two-thirds of human genes known to be involved in cancer have counterparts in the fruit fly.
-
A comparative genomics analysis of six species of yeast prompted scientists to significantly revise their initial catalog of yeast genes and to predict a new set of functional elements that play a role in regulating genome activity, not just in yeast but across many species.
-
Researchers studying milk production have mapped genes that increase the yield of high-fat milk in cows, resulting in higher production levels and potentially a significant economic impact. This is one of many studies aimed at increasing food production.
-
Scientists have found genes that increase muscling in cattle by twofold; they found the same genes in racing dogs, and such results may foster human performance studies.
-
Comparisons of nearly 50 bird species' genomes revealed a gene network that underlies singing in birds and that may have an important role in human speech and language. The bird researchers also found gene networks responsible for traits such as feathers and beaks.
-
In recent years, researchers in the National Human Genome Research Institute (NHGRI) intramural program also have studied the genomics of various cancer types in dogs, including common cancers and other diseases, to try to develop new insights into the human form of the condition. In some cases, they have mapped genes contributing to these disorders.
-
In other studies, NHGRI researchers are comparing how genes affect body shape and size in dogs to better understand growth and development. Studies of dogs with sleep problems have revealed genes and pathways - and potential drug targets - to treat sleep problems.
-
What other genomes have been sequenced?
Researchers have sequenced the complete genomes of hundreds of animals and plants-more than 250 animal species and 50 species of birds alone-and the list continues to grow almost daily.
In addition to the sequencing of the human genome, which was completed in 2003, scientists involved in the Human Genome Project sequenced the genomes of a number of model organisms that are commonly used as surrogates in studying human biology. These include the rat, puffer fish, fruit fly, sea squirt, roundworm, and the bacterium Escherichia coli. For some organisms NHGRI has sequenced many varieties, providing critical data for understanding genetic variation.
DNA sequencing centers supported by NHGRI also have sequenced genomes of the chicken, dog, honey bee, gorilla, chimpanzee, sea urchin, fungi and many other organisms.
-
What other genomes have been sequenced?
Researchers have sequenced the complete genomes of hundreds of animals and plants-more than 250 animal species and 50 species of birds alone-and the list continues to grow almost daily.
In addition to the sequencing of the human genome, which was completed in 2003, scientists involved in the Human Genome Project sequenced the genomes of a number of model organisms that are commonly used as surrogates in studying human biology. These include the rat, puffer fish, fruit fly, sea squirt, roundworm, and the bacterium Escherichia coli. For some organisms NHGRI has sequenced many varieties, providing critical data for understanding genetic variation.
DNA sequencing centers supported by NHGRI also have sequenced genomes of the chicken, dog, honey bee, gorilla, chimpanzee, sea urchin, fungi and many other organisms.
How is NHGRI involved in the growth of this new field of research?
NHGRI pioneered the development of DNA sequencing methods and technologies - including informatics - and has funded research to study the genomes of a wide range of species. The National Institutes of Health (NIH) Intramural Sequencing Center has been instrumental in the sequencing of many organisms.
NHGRI programs such as ENCODE (Encyclopedia of DNA Elements) and modENCODE (model organism Encyclopedia of DNA Elements) have compared and contrasted the inner workings of animal and human genomes to try to better understand how genomes function.
In modENCODE, researchers found shared patterns of gene activity and regulation among fly, worm and human genomes. The mouse ENCODE Consortium demonstrated that, in general, the systems that are used to control gene activity have many similarities in mice and humans.
-
How is NHGRI involved in the growth of this new field of research?
NHGRI pioneered the development of DNA sequencing methods and technologies - including informatics - and has funded research to study the genomes of a wide range of species. The National Institutes of Health (NIH) Intramural Sequencing Center has been instrumental in the sequencing of many organisms.
NHGRI programs such as ENCODE (Encyclopedia of DNA Elements) and modENCODE (model organism Encyclopedia of DNA Elements) have compared and contrasted the inner workings of animal and human genomes to try to better understand how genomes function.
In modENCODE, researchers found shared patterns of gene activity and regulation among fly, worm and human genomes. The mouse ENCODE Consortium demonstrated that, in general, the systems that are used to control gene activity have many similarities in mice and humans.
Last updated: August 15, 2020