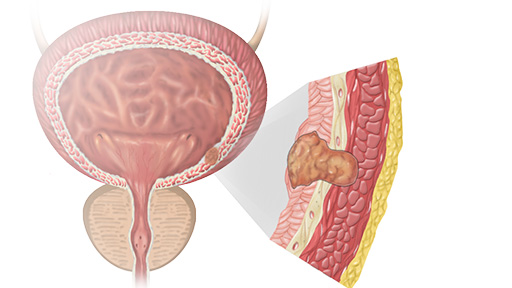
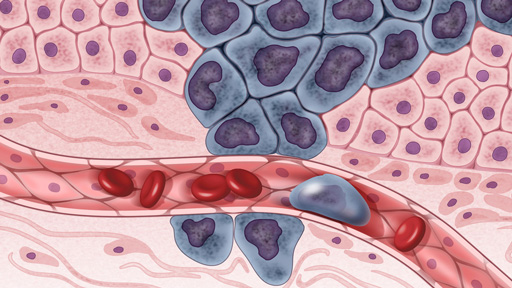
Why do some patients stay in remission, while others see their cancer return?
To get a better idea of who will be more likely to relapse, researchers are trying to understand a process whose rules are constantly being written and rewritten by the cancer's rogue cells. Untangling the genetic influences on these rules will help doctors better understand what mutations - changes to the cancer cell's DNA - may lead to relapse.
In recent research published online in July in the journal Leukemia, Paul Liu, M.D., Ph.D., asenior investigator and Raman Sood, Ph.D., an associate investigator in the Translational and Functional Genomics Branch at the National Human Genome Research Institute, and their collaborators studied 10 patients initially diagnosed with a specific kind of relapse-prone acute myeloid leukemia (AML). Though AML is generally thought to be a childhood disease, almost 21,000 American adults will be diagnosed with AML this year, according to the American Cancer Society.
"We're trying to understand why leukemia patients relapse, and if there are any DNA-level mutations that account for the leukemia coming back." said Dr. Liu. "Overall 5-year survival is about 50 percent, and a big reason is because of relapse - so it's a major issue to managing the disease."
The 10 patients in this study each saw their cancer relapse after an initial AML diagnosis and treatment. To study what may have caused the relapse specifically, bone marrow samples were taken at three specific times - diagnosis, remission and relapse - and then subjected to a battery of genetic and genomic analyses. This allowed the team to study the cancer cells' genome. A genome can be thought of as a huge DNA "book" in each cell that includes all of our genes (the chapters), and provides all the instructions for our cells to do their jobs. Because cancer cells have many mutations in their genes, there are significant differences between the genome of a healthy cell and that of a cancer cell.
The analysis of the cancer cells' genome at the three time-points is akin to reading through the chapters (genes) of the cancer's book (the genome) while it still being edited, looking page by page. The key part is spotting the edits (mutations) between each version. By comparing the results from the three sequential samples (the list of mutations/edits), the team was able to view the evolution of the mutations that occur throughout the cancer cell's genome. From there, they could tell which mutations were in the original sample, and which were in the relapse samples - possibly providing clues as to which mutation(s) spurs a relapse.
Using this time-lapse technique - a first for this type of AML - the researchers discovered new mutations occurring frequently in the relapse samples, illustrating the unstable nature of cancer cells' genomes. After sifting through the new mutations, they discovered two important genes that, when mutated, seemed to cause a relapse and drive blood cells that are still in production in the bone marrow to instead become leukemia cells.
The gene that appears to lead to relapse, GATA2, is a sort of "master" switch that controls blood cell production in the bone marrow. In other research, this gene has been implicated in causing AML, but this is the first time it has ever been linked to relapse.
The other gene that leads to leukemia cell production when mutated, DHX15, appears to be a new "driver gene" that seems to lead to instability in a key cell function that helps translate the DNA instructions so that the cell can use them. Identifying these cancer driver genes can help researchers unravel how a cancer starts, and also hopefully lead to finding new targets for treatment.
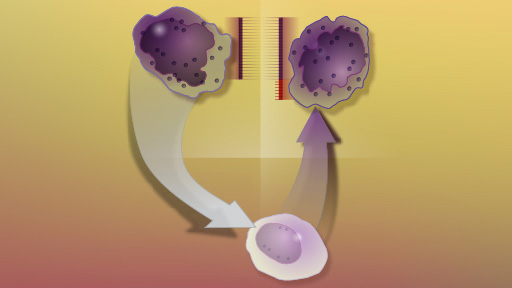
In addition to these discoveries, the time-lapse nature of the data provided insight into how relapse occurs. Drs. Liu and Sood proposed two paths: either through a pre-leukemia cell (a "sub-clone") that was unaffected by radiation or chemotherapy treatments and eventually became leukemia after accumulating more mutations; or through a few leukemia cells that had escaped treatment and returned with additional mutations. The team found signs of each path, with about half the patients showing signs that their relapsed AML was due to a sub-clone cell, and the other half showing evidence that some of the initial leukemia cells had evaded treatment.
Gaining better insight into the genetic landscape of AML can help doctors determine if a particular patient is more likely to face a relapse, while also giving pharmaceutical companies potential targets for new drugs. Further, in the future a doctor may order a genetic test on the cancer cells to search for genetic relapse markers, like a GATA2 mutation. Finding this (or another) marker, the doctor may order more frequent blood tests to monitor for signs of relapse, while also initiating the search for a compatible bone marrow donor.
Though this was a small study and the results are preliminary, the findings likely have significance beyond AML. Because cells follow certain rules outlined by genes, other cancers may relapse in much the same way. Identifying AML's genetic mutations and genetic pathways of relapse may help all cancer researchers look at these same genes (or others) to determine if they see a similar pattern.
Read the Study
Sood, R., Hansen, N., Donovan, F., Carrington, B., Bucci, D., Maskeri, B., ... Liu, P. Somatic mutational landscape of AML with inv(16) or t(8;21) identifies patterns of clonal evolution in relapse leukemia. Leukemia, doi: 10.1038/leu.2015.141. 2015. [Full Text]