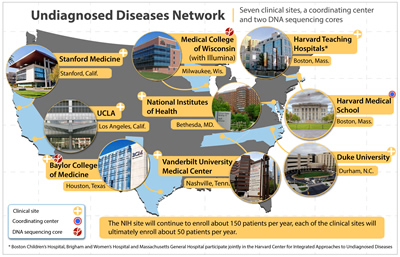
The two universities will become the DNA Sequencing Core sites for the UDN. The UDN also includes seven clinical sites at academic medical centers and at the NIH, as well as a coordinating center operated by Harvard Medical School in Boston.
In addition, NIH announced six UDN grants that will fund studies of the function of genes thought to play roles in particular patients' conditions.
Christine M. Eng, M.D., professor of molecular and human genetics and medical director of the DNA Diagnostic Laboratory, is principal investigator at Baylor College of Medicine's UDN DNA Sequencing Core. The Baylor College of Medicine facility will provide sequencing of the protein-coding portion of the genome, or exome. The clinical sequencing results will meet standards set by the Clinical Laboratory Improvement Amendments (CLIA), which govern lab tests for diagnostic, treatment of health assessment or prevention.
Howard J. Jacob, Ph.D., professor of physiology and human and molecular genetics and director of the Human and Molecular Genetics Center, is principal investigator at the Medical College of Wisconsin's UDN DNA Sequencing Core. The facility will provide CLIA-certified genome sequence data for UDN patients and their family members in partnership with the sequencing and array technology company, Illumina.
The UDN, based on a pilot program launched in 2008 at the NIH Clinical Center in Bethesda, Maryland, is developing approaches for the diagnosis of difficult-to-solve medical cases. Along with traditional methods, doctors pursue medical diagnoses for patients by analyzing DNA sequence, which can reveal variants in patients' genome that may be the cause of a disease. Genome sequencing is an increasingly fast and inexpensive technology that only recently has been used for patient diagnosis.
Genome sequencing for patients from the NIH Undiagnosed Diseases Program (UDP) has identified individual variation in genes that clinical researchers want to understand better, especially how their function or malfunction may affect a disease condition. Investigation of gene function is a critical step in the process leading to diagnoses and potential treatments for patients with these rare or new diseases.
Recipients of the new NIH awards to study gene function (pending available funds) are:
- Baylor College of Medicine, $470,000 over two years
Principal Investigator: Brett H. Graham, M.D.
The study will entail cell culture and fruit fly experiments through which researchers will develop a pipeline of models for disease-causing mutations in a broad range of human genes.
- Johns Hopkins University, $486,000 over two years
Principal Investigator: Paul F. Worley, M.D.
Researchers will characterize the function of a mutation in the FRMPD4/Preso1 gene that is thought to be responsible for neurological symptoms in two UDP patients. The investigation will use a mouse model of the gene mutation as well as cell culture studies.
- University of California Los Angeles, $462,000 over two years
Principal Investigator: Shuo Lin, Ph.D.
By developing a zebrafish model of undiagnosed disease genes, researchers will provide a system through which such genes can be functionally characterized. The researchers will make observations of the outcome in zebrafish from inserting mutations in genes that have a parallel with human disease.
- University of Iowa, $340,000 over two years
Principal Investigator: Diane Slusarski, Ph.D.
Researchers will use a zebrafish model to explore the function of 15 gene mutations associated with epilepsy.
- University of Michigan, $467,000 over two years
Principal Investigator: Anthony Antonellis, Ph.D.
Using a zebrafish model, researchers will evaluate the role that six gene variants play in loss of function of the gene that codes for the aminoacyl-tRNA synthetase enzyme. Changes in these genes have been identified in patients in the UDP Program.
- Vanderbilt University, $432,000 over two years
Principal Investigator: Wenbiao Chen, Ph.D.
Researchers will establish a pipeline for functional analysis, drug discovery, and pathway identification of UDP-identified genes that can be studied in zebrafish.
The UDN is funded through the NIH Common Fund, which supports cross-cutting programs that are expected to have exceptionally high impact. The goals of the program are to investigate compelling research questions that the study of undiagnosed conditions entails, to identify new disorders or rare presentations of known diseases and to shed light on the disease causes and potential targets for treatments.
For more information about the UDN, visit http://commonfund.nih.gov/Diseases/index.




