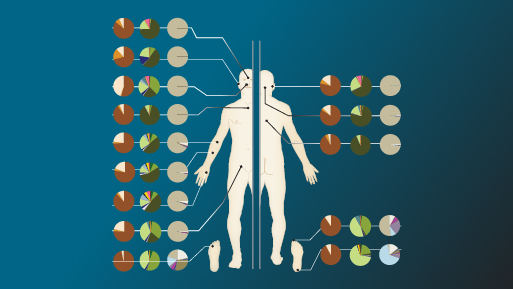
The human microbiome comes into focus
Not so long ago, researchers thought that if they could identify the tens of thousands of microbial species that live in or on human beings - the human microbiome - they would find therapies for a host of diseases. As it turned out, the problem isn't that simple: the key question isn't who our fellow travelers are but how they interact with our bodies.
Exploring the Host-Microbiome Ecyostem
Now, in the second phase of the Human Microbiome Project, three research teams funded by the National Institutes of Health (NIH) are gathering data at the most basic molecular levels about the biological processes that make up the host-microbiome ecosystem. The three conditions chosen as test beds - because of their wide impact and because they can serve as models of other microbiome-associated human conditions - are pregnancy and preterm birth, onset of inflammatory bowel disease and onset of type 2 diabetes.
Importantly, as with many projects in the genomics research community, the researchers are committed to releasing their data rapidly and comprehensively, so that other scientists can use the results for their own research into the role of the microbiome in specific diseases.
In the September 10, 2014 issue of Cell Host & Microbe, the Integrative Human Microbiome Project Research Consortium - as the current consortium is known - detailed what data are being collected and into which databases the results will be deposited.
"We're producing an incredibly rich array of data for the community from the microbiomes and hosts in these cohorts, so that scientists can evaluate for themselves with these freely available data which properties are the most relevant for understanding the role of the microbiome in the human host," said Lita M. Proctor, Ph.D., program director of the Human Microbiome Project at NIH's National Human Genome Research Institute (NHGRI).
"The members of the Consortium can take advantage of each other's expertise in dealing with some very complex science in these projects," she said. "We're generating these data as a community resource and we want to describe this resource in enough detail so people can anticipate the data that will be produced, where they can find it and the analyses that will come out of the Consortium's efforts."
NHGRI Director Eric D. Green, M.D., Ph.D., said the data will help scientists "wrangle the almost overwhelming numbers of features of the microbiome into a manageable subset of features, in order to evaluate which properties are the most informative for deciphering the role of the microbiome in human health and disease."
Using cutting-edge DNA sequencing technology, mass spectroscopy and other 'omics technologies, researchers are learning much about the microbiome's complexities. In the first phase of the Human Microbiome Project, which ended in 2012, researchers detected the presence of more than 10,000 species of bacteria living on or within human beings and aiding their hosts in functions such as digestion, energy production, protection against invading pathogens and immune system support. Now, the number of bacterial species living with human beings is thought to be closer to 15,000.
Other estimates have changed, too. For years the number of microbial cells on or in each human was thought to outnumber human cells by 10 to 1. This now seems a huge understatement. Dr. Proctor noted that the 10-to-1 estimate was based only on bacterial cells, but the microbiome also includes viruses, protozoa, fungi and other forms of microscopic life. "So if you really look at the entire microbial community, you're probably looking at more like a 100-to-1 ratio," she said.
Although thousands of bacterial species may make their homes with human beings, each individual person is host to only about 1,000 species at a time, according to the findings of the Human Microbiome Project's first phase in 2012.
In addition, judging from the array of common functions of bacterial genes, if the bacteria are healthy, each individual's particular suite of species appear to come together to perform roughly the same biological functions as another healthy individual. In fact, researchers found that certain bacterial metabolic pathways were always present in healthy people, and that many of those pathways were often lost or altered in people who were ill.
When Phase 2 of the Human Microbiome Project began in 2013, Dr. Proctor said, "We decided to fund three well-characterized cohort studies that were good models for microbiome-associated conditions, and to collect all the properties we could, not only of the microbiome but of the host. We did not want to go back and wish we had collected more information from these human microbiome model studies."
The three-year studies, which have received $24.2 million to date, include:
- Pregnancy and Preterm Birth. The "Multi-omic Microbiome study: Pregnancy Initiative" will study changes in the vaginal microbiome and host responses during pregnancy, including preterm birth, labor and delivery, as well as acquisition of microbiomes by newborns. About 2,000 women will be followed through their pregnancies and deliveries.
Preterm birth is the leading cause of neonatal morbidity and death worldwide, and in the United States, the cost of health care for newborns with complications totals more than $26 billion a year. The maternal microbiome is known to undergo significant changes during pregnancy, and researchers suspect that these changes play an important role in the health of the woman and her fetus. This role currently is poorly understood.
Researchers led by Gregory A. Buck, Ph.D., Kimberly K. Jefferson, Ph.D., and Jerome F. Strauss III, M.D., Ph.D., of Virginia Commonwealth University in Richmond will analyze samples from various body sites using the full panoply of existing genomic, proteomic, metabolic, transcriptomic and other molecular tests - collectively called "multi-omic" - to try to work out just what role the microbiome plays in birth and neonatal death.
- Onset of Inflammatory Bowel Disease. Inflammatory bowel disease (IBD), which affects nearly 1.5 million Americans, is believed to be closely linked to microorganisms in the gut, but no specific microbe or population of microbes has been identified as the infectious agents. This project will attempt to pry apart the entire gut microbial ecosystem to determine precisely which microbes are interacting with which microbes and with their human host.
Ramnik J. Xavier, M.D., Ph.D., of Massachusetts General Hospital and Harvard Medical School, both in Boston, and the Broad Institute of MIT and Harvard in Cambridge, Massachusetts, and Curtis Huttenhower, Ph.D., of the Harvard School of Public Health in Boston and the Broad Institute, and their co-workers will follow four groups of volunteers for a year: new-onset pediatric patients, new-onset adult patients, people with Crohn's disease or ulcerative colitis and a control group of healthy people. The scientists hope to uncover the molecular mechanisms by which the gut microbiome causes disease and to determine whether identifying the specific microbial population can guide therapy.
In addition, the project aims to build a new IBD database containing multi-omic data -that is, all the molecular information that is possible to gather using current technology - as a resource for other scientists.
- Type 2 Diabetes Onset. To explore the relationship between the microbiome, stress and the onset of type 2 diabetes, the researchers will follow about 60 people at risk for the disease for three years. They will analyze microbiome and host samples from multiple sites every two to three months and more frequently during times of stress from respiratory illness or other causes.
The scientists - led by Michael Snyder, Ph.D., of Stanford University in Stanford, California, and George Weinstock of the Jackson Laboratory for Genomic Medicine in Farmington, Connecticut - believe that immunological stressors such as the flu, as well as environmental or emotional stress also - cause changes in specific biological pathways. These in turn can cause changes in the microbiome, and possibly trigger onset of disease linked to the immune system. Such stressors could particularly affect individuals prone to develop autoimmune disease.
They expect to find changes in both microbiome and host "at an unprecedented level and frequency" when the volunteers are under stress, and they will try to identify the molecules and pathways involved through multi-omic analyses. Their findings could be relevant not just to type 2 diabetes - which now affects about 21 million Americans and is expected to double by 2050 - but also to other autoimmune diseases.
-
Exploring the Host-Microbiome Ecyostem
Now, in the second phase of the Human Microbiome Project, three research teams funded by the National Institutes of Health (NIH) are gathering data at the most basic molecular levels about the biological processes that make up the host-microbiome ecosystem. The three conditions chosen as test beds - because of their wide impact and because they can serve as models of other microbiome-associated human conditions - are pregnancy and preterm birth, onset of inflammatory bowel disease and onset of type 2 diabetes.
Importantly, as with many projects in the genomics research community, the researchers are committed to releasing their data rapidly and comprehensively, so that other scientists can use the results for their own research into the role of the microbiome in specific diseases.
In the September 10, 2014 issue of Cell Host & Microbe, the Integrative Human Microbiome Project Research Consortium - as the current consortium is known - detailed what data are being collected and into which databases the results will be deposited.
"We're producing an incredibly rich array of data for the community from the microbiomes and hosts in these cohorts, so that scientists can evaluate for themselves with these freely available data which properties are the most relevant for understanding the role of the microbiome in the human host," said Lita M. Proctor, Ph.D., program director of the Human Microbiome Project at NIH's National Human Genome Research Institute (NHGRI).
"The members of the Consortium can take advantage of each other's expertise in dealing with some very complex science in these projects," she said. "We're generating these data as a community resource and we want to describe this resource in enough detail so people can anticipate the data that will be produced, where they can find it and the analyses that will come out of the Consortium's efforts."
NHGRI Director Eric D. Green, M.D., Ph.D., said the data will help scientists "wrangle the almost overwhelming numbers of features of the microbiome into a manageable subset of features, in order to evaluate which properties are the most informative for deciphering the role of the microbiome in human health and disease."
Using cutting-edge DNA sequencing technology, mass spectroscopy and other 'omics technologies, researchers are learning much about the microbiome's complexities. In the first phase of the Human Microbiome Project, which ended in 2012, researchers detected the presence of more than 10,000 species of bacteria living on or within human beings and aiding their hosts in functions such as digestion, energy production, protection against invading pathogens and immune system support. Now, the number of bacterial species living with human beings is thought to be closer to 15,000.
Other estimates have changed, too. For years the number of microbial cells on or in each human was thought to outnumber human cells by 10 to 1. This now seems a huge understatement. Dr. Proctor noted that the 10-to-1 estimate was based only on bacterial cells, but the microbiome also includes viruses, protozoa, fungi and other forms of microscopic life. "So if you really look at the entire microbial community, you're probably looking at more like a 100-to-1 ratio," she said.
Although thousands of bacterial species may make their homes with human beings, each individual person is host to only about 1,000 species at a time, according to the findings of the Human Microbiome Project's first phase in 2012.
In addition, judging from the array of common functions of bacterial genes, if the bacteria are healthy, each individual's particular suite of species appear to come together to perform roughly the same biological functions as another healthy individual. In fact, researchers found that certain bacterial metabolic pathways were always present in healthy people, and that many of those pathways were often lost or altered in people who were ill.
When Phase 2 of the Human Microbiome Project began in 2013, Dr. Proctor said, "We decided to fund three well-characterized cohort studies that were good models for microbiome-associated conditions, and to collect all the properties we could, not only of the microbiome but of the host. We did not want to go back and wish we had collected more information from these human microbiome model studies."
The three-year studies, which have received $24.2 million to date, include:
- Pregnancy and Preterm Birth. The "Multi-omic Microbiome study: Pregnancy Initiative" will study changes in the vaginal microbiome and host responses during pregnancy, including preterm birth, labor and delivery, as well as acquisition of microbiomes by newborns. About 2,000 women will be followed through their pregnancies and deliveries.
Preterm birth is the leading cause of neonatal morbidity and death worldwide, and in the United States, the cost of health care for newborns with complications totals more than $26 billion a year. The maternal microbiome is known to undergo significant changes during pregnancy, and researchers suspect that these changes play an important role in the health of the woman and her fetus. This role currently is poorly understood.
Researchers led by Gregory A. Buck, Ph.D., Kimberly K. Jefferson, Ph.D., and Jerome F. Strauss III, M.D., Ph.D., of Virginia Commonwealth University in Richmond will analyze samples from various body sites using the full panoply of existing genomic, proteomic, metabolic, transcriptomic and other molecular tests - collectively called "multi-omic" - to try to work out just what role the microbiome plays in birth and neonatal death.
- Onset of Inflammatory Bowel Disease. Inflammatory bowel disease (IBD), which affects nearly 1.5 million Americans, is believed to be closely linked to microorganisms in the gut, but no specific microbe or population of microbes has been identified as the infectious agents. This project will attempt to pry apart the entire gut microbial ecosystem to determine precisely which microbes are interacting with which microbes and with their human host.
Ramnik J. Xavier, M.D., Ph.D., of Massachusetts General Hospital and Harvard Medical School, both in Boston, and the Broad Institute of MIT and Harvard in Cambridge, Massachusetts, and Curtis Huttenhower, Ph.D., of the Harvard School of Public Health in Boston and the Broad Institute, and their co-workers will follow four groups of volunteers for a year: new-onset pediatric patients, new-onset adult patients, people with Crohn's disease or ulcerative colitis and a control group of healthy people. The scientists hope to uncover the molecular mechanisms by which the gut microbiome causes disease and to determine whether identifying the specific microbial population can guide therapy.
In addition, the project aims to build a new IBD database containing multi-omic data -that is, all the molecular information that is possible to gather using current technology - as a resource for other scientists.
- Type 2 Diabetes Onset. To explore the relationship between the microbiome, stress and the onset of type 2 diabetes, the researchers will follow about 60 people at risk for the disease for three years. They will analyze microbiome and host samples from multiple sites every two to three months and more frequently during times of stress from respiratory illness or other causes.
The scientists - led by Michael Snyder, Ph.D., of Stanford University in Stanford, California, and George Weinstock of the Jackson Laboratory for Genomic Medicine in Farmington, Connecticut - believe that immunological stressors such as the flu, as well as environmental or emotional stress also - cause changes in specific biological pathways. These in turn can cause changes in the microbiome, and possibly trigger onset of disease linked to the immune system. Such stressors could particularly affect individuals prone to develop autoimmune disease.
They expect to find changes in both microbiome and host "at an unprecedented level and frequency" when the volunteers are under stress, and they will try to identify the molecules and pathways involved through multi-omic analyses. Their findings could be relevant not just to type 2 diabetes - which now affects about 21 million Americans and is expected to double by 2050 - but also to other autoimmune diseases.
- Pregnancy and Preterm Birth. The "Multi-omic Microbiome study: Pregnancy Initiative" will study changes in the vaginal microbiome and host responses during pregnancy, including preterm birth, labor and delivery, as well as acquisition of microbiomes by newborns. About 2,000 women will be followed through their pregnancies and deliveries.
Related Content
Last updated: September 10, 2014